When it comes to massively parallel sequencing, few areas of human health stand to benefit as much as rare genetic diseases. Indeed, both whole-genome and exome sequencing strategies have identified disease-causing mutations in probands with Charcot-Marie Tooth disease, Miller syndrome, severe brain malformations, and a few other disorders. The Mito10K project took a different approach. They assembled a cohort of mostly unrelated individuals with complex I deficiency (n=103), the most common cause of human respiratory chain diseases.
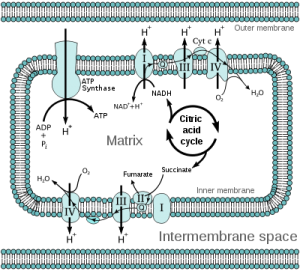
Mitochondrial Electron Transport Chain (Wikipedia)
Forty-two HapMap samples were included as controls. Instead of employing a whole-genome or exome strategy, they performed deep resequencing of carefully-chosen candidate genes in pools of ~20 samples. And they did it all using a single Illumina flowcell.
Pooled Sequencing of Candidate Genes
The candidates included 103 genes that (i) encoded known complex I proteins, (ii) were implicated in the disease, or (iii) were identified by phylogenetic profiling. The 145 kb target space comprised 653 exons from nuclear genes (138 kb) and two mtDNA regions (7 kb). About 90% of target regions achieved at least 100x coverage; the median redundancy was 3,359x per pool, which works out to ~168x per individual. Next, the authors developed a method (“Syzygy”) to model sequencing error and call variants at very low frequencies. A comparison of calls for the HapMap samples to existing genotype data suggested 92% sensitivity and 99.6% specificity, at sites where coverage was 100x or greater.
Although the pooling strategy worked well for nuclear DNA, there were some problems with the targeted regions in mtDNA. Basically, the distribution of mtDNA was not uniform between samples. That may be due to the fact that while each cell contains exactly 2 copies of each nuclear chromosome, it contains numerous mitochondria and thus numerous copies of the MT chromosome (possibly 20-25 per cell, by one estimate). The resulting shift in sample representation can be quite dramatic. In one pool, for example, 96% of the mtDNA came from a single individual (5% of the pool). The bottom line is that sensitivity to call mutations in pooled samples is going to be lower for mtDNA.
Variant Calling and “Deleteriousness” Prioritization
The unfortunately-named Syzygy method identified 652 variants (high confidence); to boost sensitivity, the authors also employed an ad-hoc approach that called 246 more variants supported by at least 3 reads on each strand (low confidence). The 898 calls were filtered to prioritize variants that seemed likely to underlie a rare and devastating phenotype. In short, the authors removed:
- Variants present in healthy individuals (HapMap controls) or public databases (dbSNP, mtDB, 1000 Genomes).
- Synonymous or noncoding variants, unless they affected tRNA or splice sites.
- Missense variants at positions of low evolutionary conservation
Of 898 detected variants, 216 remained and were validated by multiplexed Sequenom genotyping. Some 82 sites were also Sanger-sequenced to assess the accuracy of the genotyping platform. The comparison revealed 11% false positives and 2% het/hom miscalls, for an overall error rate of 13% for Sequenom assays. Ouch. As for the variant calls, the validation rate was pretty good for high-confidence calls (91/109, or 84%) but rather abysmal for the low-confidence ones (12/107, or 11%). Intriguingly, validation assays identified 12 additional pathogenic variants that were missed by the discovery screen. Based on these data, the sensitivity of the Syzygy method alone was 79.1% (91/115). That’s not bad, but probably not enough for a study whose goal is to identify rare disease-causing variants.
New Diagnoses from Validated Mutations
Some 60 of the sequenced cases lacked a previous molecular-genetic diagnosis. Among these, the authors were able to provide 11 new diagnoses based on mutations in known disease-causing genes. Several lines of supporting evidence were given to support the diagnoses:
- 6 patients had mutations that were previously known to be disease-causing.
- 3 patients were homozygous for deleterious mutations that caused splicing defects (observed in cDNA) and no detectable protein (by SDS-page and protein blot).
- 2 patients had mutations in highly conserved protein domains.
Intriguingly, half of the cases with known mutations (3/6) were compound heterozygotes; that is, they inherited a different defect in the same gene from mother and father. This apparent prevalence of compound hets in monogenic disease is unsettling because they tend to make pedigree analysis complicated and require detection of both variants in heterozygous form, which is more difficult to do by sequencing.
Detection and Characterization of Novel Disease Genes
The key finding of this paper (as suggested by the title) was the implication of two new genes in complex I deficiency: NUBPL and FOXRED1. Pathogenicity of each mutated genes was confirmed by a “rescue” assay in which introduction of wild-type cDNA into patient fibroblasts restored complex I activity. In the absence of rescue, residual complex I activity was markedly reduced (19-40%) in the NUBPL-mutated fibroblasts and strikingly reduced (9-15%) in the FOXRED1-mutated fibroblasts.
The case with NUBPL mutations was particularly interesting. RT-PCR showed that the dominant mRNA species was truncated, and the full-length transcript hardly expressed at all. Sequencing revealed that the shortened fragment had a branch site mutatation that likely caused exon 10 skipping, as well as a missense mutation (Gly56Arg), both on the paternal chromosome. The maternal allele wasn’t expressed. Array-based copy number analysis, however, showed that the maternal chromosome had a complex rearrangement of NUBPL in which exons 1-4 were deleted and exon 7 was duplicated. Obviously this structural variation was not detected in the discovery screen. I think this highlights two things: the importance of structural variation in human disease, and the limitations of targeted sequencing on NGS platforms.
Success and Limitations
As the authors note in their discussion, key to the success of this study was the availability of cellular models of disease, with which the pathogenicity of newly discovered mutations in individual patients could be established. With the two new findings, the 11 newly diagnosed cases, and the 40 or so already-diagnosed cases, the authors now have identified the genetic defect for about half of the cases in their cohort. What about the rest? The authors admit that the causal mutations were likely missed because:
- They occur in genes not targeted in this study
- They affect targeted genes, but reside in noncoding regulatory regions or novel/unknown exons
- They were targeted, but not detected due to limited sensitivity (especially in mtDNA)
- They were detected, but filtered out as not likely to be deleterious
- They are large-scale deletions or rearrangements, which this approach can’t detect
Despite these limitations, the authors have demonstrated that sequencing carefully-chosen candidate genes in pooled samples, with follow-up validation and experimental support, can successfully identify disease-causing mutations in a good-sized patient cohort. Not bad for a single flowcell.
References
Calvo, S., Tucker, E., Compton, A., Kirby, D., Crawford, G., Burtt, N., Rivas, M., Guiducci, C., Bruno, D., Goldberger, O., Redman, M., Wiltshire, E., Wilson, C., Altshuler, D., Gabriel, S., Daly, M., Thorburn, D., & Mootha, V. (2010). High-throughput, pooled sequencing identifies mutations in NUBPL and FOXRED1 in human complex I deficiency Nature Genetics, 42 (10), 851-858 DOI: 10.1038/ng.659
Ng SB, Buckingham KJ, Lee C, et al (2010). Exome sequencing identifies the cause of a mendelian disorder. Nature genetics, 42 (1), 30-5 PMID: 19915526
Bilgüvar K, Oztürk AK, Louvi A, et al (2010). Whole-exome sequencing identifies recessive WDR62 mutations in severe brain malformations. Nature, 467 (7312), 207-10 PMID: 20729831
Lupski JR, Reid JG, Gonzaga-Jauregui C, et al (2010). Whole-genome sequencing in a patient with Charcot-Marie-Tooth neuropathy. The New England journal of medicine, 362 (13), 1181-91 PMID: 20220177
Lalonde E, Albrecht S, Ha KC, et al (2010). Unexpected allelic heterogeneity and spectrum of mutations in Fowler syndrome revealed by next-generation exome sequencing. Human mutation, 31 (8), 918-23 PMID: 20518025