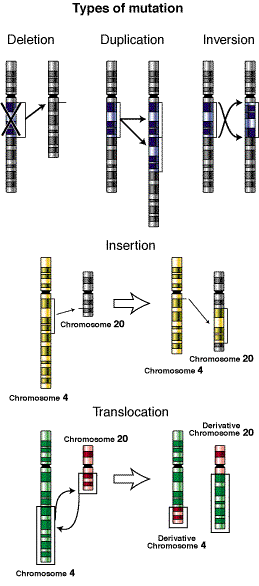
Two recent papers used very different appraoches to shed light on the genetic alterations underlying tumor growth and progression in human cancers. Peter Campbell and colleagues from the Wellcome Trust Sanger Institute employed Illumina paired-end sequencing to survey the landscape of structural variation in metastatic pancreatic cancer. Ivana Bozic and colleagues from Harvard University took a different approach – they constructed mathematical models of tumor progression via the accumulation of driver and passenger mutations. I happened to read both papers on a long airplane ride, and learned a great deal about mutations and metastasis in human cancers.
Pancreatic Cancer: Bad News
You learn a lot from the introduction sections of these papers, even if the Letter to Nature format keeps them short. I knew that pancreatic cancer had, in general, a poor prognosis. It turns out that the five year mortality for this cancer is 97-98%, usually due to “widespread metastatic disease.” These tumors also appear to carry a heavy mutational load. A 2008 survey of 24 pancreatic cancers (by Bert Vogelstein’s group at Johns Hopkins) found that tumors had ~63 genetic alterations on average, the majority of which were point mutations. Copy number changes are also common in this cancer type. Frequently mutated genes include tumor suppressors (TP53, SMAD4, CDKN2A) as well as oncogenes (KRAS, MYC). Less was known about the patterns of structural variation in pancreatic cancer.
Detecting Rearrangements by Paired-End Sequencing
Peter Campbell’s group has developed a very nice strategy for identifying somatically acquired rearrangments by massively parallel paired-end sequencing on the Illumina platform. They’ve already applied it to the characterization of SVs in several cancer cell lines. In this study, they generated 50-150 million read pairs (2 x 37 bp) per patient, which, in their experience, enables detection of 50-60% of rearrangements in a sample. Across the 13 pancreatic tumors, they identified 381 somatic and 177 germline rearrangements across seven categories: amplicon, deletion, tandem duplication, inversion, fold-back inversion, interchromosomal (translocation), and “other” intrachromosomal.
Many rearrangements corresponded with a change in copy number. In one metastasis, for example, numerous rearrangements (some inverted, some not) combine to amplify the KRAS oncogene.
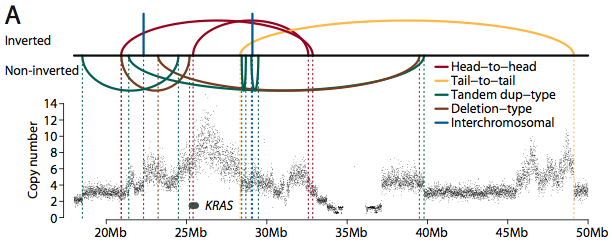
Rearrangement/Amplification of KRAS (Credit: Nature).
Fold-back Inversions and Inter-Lesion Genetic Heterogeneity
One sixth of the rearrangements identified fell into a class the authors call “fold-back” inversions. These are genomic regions that are duplicated, but the two copies face in opposite directions from the breakpoint (as opposed to a tandem duplication). The authors suggest breakage-fusion-bridge cycles as the likely mechanism that creates such an event. Basically, a double-stranded break that occurs during G0-G1 phase is replicated (in S phase), creating two duplicated end sequences. These are fused together by DNA repair processes, resulting in a sort of inverted duplication (fold-back inversion) with two centromeres. These “dicentric” chromosomes are unstable, and frequently initiate the amplification of oncogenes.
Each rearrangement was [laboriously] genotyped by PCR in both the index tumor sample and matched normal control to verify the somatic status. Further, PCR and capillary sequencing were employed to resolve breakpoints, and some 206 rearrangements were genotyped across multiple lesions (metastases) in the 10 patients for which metastatic samples were available. There was a considerable amount of genetic heterogeneity among samples from the same patient. While the majority of rearrangements were present in all samples but not the germline (omnipresent); several were present in some samples but not others (partially shared) or unique to the index tumor sample (private).
Telomere Loss and Breakpoint-Fusion-Bridge Cycles
Fold-back inversions were significantly more likely than other classes of rearrangement to be omnipresent, suggesting that they occur early during tumor progression, before cancer cells disseminate. Because breakage-fusion-bridge cycles are often initiated by telomere loss, the activity of telomerase to maintain telomeres may play a pivotal role in the development of pancreatic cancer. Other studies have shown that telomerase expression is low in early tumor stages, but markedly increased in the invasive tumor. The increased expression likely suppresses breakage-fusion-bridge cycles, which may help explain why fold-back inversions are more likely to occur earlier in the development of the disease.
Ongoing Evolution in Tumors and Mets
In several patients, the authors found rearrangements that were in the primary tumor and some metastases, but not all of them. The most likely explanation for such a pattern is that the metastases were “seeded” by different cells from the primary tumor. This is intriguing, because it suggests ongoing clonal evolution, in the primary tumor, among cells capable of initiating metastases. There were also rearrangements in some metastases that weren’t detected in the primary tumor, suggesting that secondary lesions, too, are undergoing clonal evolution.
Overall, the authors demonstrated that pancreatic cancers and secondary invasions show a substantial amount of genetic heterogeneity within the same patient. There’s certainly more to be done to get the full picture of genetic alterations in these tumors, but at just ~4-10 Gbp of data per sample, the scope and nature of what the authors have uncovered is pretty impressive.
Drivers and Passengers
The other paper (contributed by Bert Vogelstein to PNAS) took a theoretical approach to modeling the accumulation of driver and passenger mutations during tumor progression. In contrast to previous models that account for only 1-2 mutations, the authors develop a model in which mutations occur sequentially in tumor cells, with each new driver mutation conferring a slightly faster growth rate. This more closely reflects recently-characterized solid tumors, which harbor 40-100 coding gene alterations, of which 5-15 are considered “driver” mutations.
Based on the assumption that any human cell contains 286 tumor suppressor genes and 91 oncogenes, the authors estimate that ~34,000 positions in the human genome could host a driver mutation. By this estimate, the driver mutation rate is approximately 3.4 x 10-5 per cell division. Under the authors’ assumption that each driver speeds tumor growth, the rate at which drivers accumulate becomes faster and faster, because the more drivers a cell has, the faster it divides. Not all mutations are successful, because they only reduce the probability that a cell will senesce or die (they don’t guarantee it). The authors considered a mutation in a tumor suppressor gene to be the central rate-limiting factor, since the other working copy tends to be lost relatively quickly due to large-scale LOH events.
Six simulated patients were modeled and presented in this study. All of them started with one driver mutation. Strikingly, though all of the input values (mutation rate, division rate) were the same, there was enormous variation in the rates of tumor progression between simulated patients. Patient 1, for example, went 20 years before acquiring a second driver mutation, and the size of the tumor remained small (<5 g). In contrast, patient 6 had a secondary driver mutation in less than 5 years; by the end of the simulation, that tumor weighed hundreds of grams. While this model is undoubtedly an oversimplification, it does highlight the importance of, well, random chance. Given the large size of the human genome and the relatively small number of potential driver mutations, an individual’s fate hinges on stochastic processes. If you’re lucky, you go decades without picking up that crucial second hit. If you’re unlucky, you don’t.
Intuitively, this seems reasonable, given the anecdotal evidence of de novo cancers, which seem to strike somewhat randomly. Of course, the older you are, the more times your cells divide, and the better chance you have of picking up additional driver mutations. And environmental exposures (like smoking and radiation exposure) certainly have a role to play, because they increase cellular mutation rates. Even so, if you believe in the model, chance plays a significant role.
Here’s to hoping you’re one of the lucky ones.
References
Bozic I, Antal T, Ohtsuki H, Carter H, Kim D, Chen S, Karchin R, Kinzler KW, Vogelstein B, & Nowak MA (2010). Accumulation of driver and passenger mutations during tumor progression. Proceedings of the National Academy of Sciences of the United States of America, 107 (43), 18545-50 PMID: 20876136
Campbell PJ, Yachida S, Mudie LJ, Stephens PJ, Pleasance ED, Stebbings LA, Morsberger LA, Latimer C, McLaren S, Lin ML, McBride DJ, Varela I, Nik-Zainal SA, Leroy C, Jia M, Menzies A, Butler AP, Teague JW, Griffin CA, Burton J, Swerdlow H, Quail MA, Stratton MR, Iacobuzio-Donahue C, & Futreal PA (2010). The patterns and dynamics of genomic instability in metastatic pancreatic cancer. Nature, 467 (7319), 1109-13 PMID: 20981101