Today our group published the second cancer genome, AML2, in the New England Journal of Medicine. In this study, we sequenced the complete genomes of tumor cells and matched normal (skin) cells from a patient with cytogenetically normal de novo FAB M1 AML. This is an exciting publication for many reasons, the foremost of which may be the venue: with an impact factor of 52.59, the NEJM is almost certainly the most widely read biomedical journal in the world.
Diagnosed with Leukemia: It Could Happen to You
The story begins three years ago, with a previously healthy 38-year-old man of European ancestry who went to his doctor complaining of fatigue and a persistent cough. After an elevated white blood cell count, his physician ordered a bone marrow biopsy, which revealed 90% cellularity and 86% blasts. Diagnosis: Leukemia.
The patient underwent ten days of chemotherapy with cytarabine (7 days) followed by daunorubicin (3 days). Five weeks later he’d obtained morphologically complete remission and recovered counts. Now, three years later, he remains in complete remission. According to my conversations with an oncologist, this kind of happy ending is not very common with leukemia. Most leukemia patients are diagnosed at an advanced age, and don’t do as well.
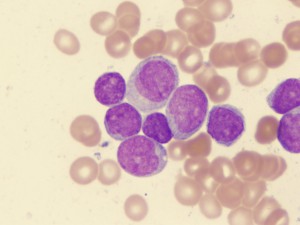
Acute myelogenous leukemia cells. Credit: Univ. of Virginia
Moving Beyond Cytogenetics
At the time of his diagnosis, routine cytogenetic analysis of the patient’s tumor cells showed a normal 46XY karyotype. Bone marrow and skin samples were banked with informed consent for whole genome sequencing in accordance with our IRB. There was no family history of leukemia, though the patient’s mother had developed breast cancer and later non-Hodgkins lymphoma. Her half sister had also developed breast cancer. The field for discovery of mutations underlying this AML was wide open.
Whole Genome Sequencing with Illumina
We sequenced the genomes of tumor cells and matched normal (skin) cells to high depth (23.3x and 21.3x, respectively) on the Illumina/Solexa platform. The tumor sample required just 16.5 runs (most of which were 2×75 PE) to reach 98% diploid coverage. That’s a dramatic improvement over our first cancer genome, AML1, which took 98 runs (36 bp SE) to achieve 91% diploid coverage. At current rates, we really can sequence a genome a week. As any bioinformatician knows, however, the analysis usually takes a bit longer.
Dave Larson in my group really deserves the credit for the whole genome variant detection pipeline applied to AML2. With direction from Elaine Mardis, Rick Wilson, and Tim Ley, and others, Dave created a pipeline for automated variant calling, somatic scoring, and tiered classification of variants for cancer genomes (see Figure 1 of the paper). We identified 3.87 million single nucleotide variants (SNVs) in the tumor genome, of which 97.5% were in the skin genome and another 1.7% were previously described (i.e. dbSNP). That left 20,256 putative somatic variants which we classified as follows:
- Tier 1 variants were coding variants that alter amino acid sequences, like nonsynonymous, nonstop, and splice-site mutations.
- Tier 2 variants were variants in evolutionarily conserved or regulatory-potential sequences of the genome.
- Tier 3 were the remaining variants that were in non-repetitive regions of the genome.
- Tier 4 were the remaining variants that were in repetitive regions of the genome.
Validation and Deep 454 Read Counts
We used 3730 sequencing to validate somatic variants in Tiers 1 and 2. Some 62 mutations were validated, of which 10 were tier 1 (amino acid-altering) mutations. Additionally, we validated two somatic indels, one of which (NPM1) was previously described; the other was an insertion in the CEP170 gene predicted to add a leucine residue to the encoded protein.
In the absence of true functional validation, there are at least two approaches to evaluating whether or not a somatic mutation is a driver – a mutation that confers some advantage to drive tumor development – or a passenger – a background mutation that’s just along for the ride. First, driver mutations should be present most tumor cells, since the dominant clone will be the most “fit” in the tumor population. To assess mutation frequencies in our patient’s tumor cells, we applied 454 sequencing of mutation-containing amplicons in the tumor DNA, tumor cDNA, and skin DNA. Deep read counts for somatic events on the X and Y chromosomes showed allele frequencies of around 98%, consistent with the fact that nearly all cells in the bone marrow sample were part of the malignant clone. For the rest of the somatic mutations, variant frequencies hovered near the 50% mark (as expected) with a few exceptions. The CEP170 indel had a reduced (~35%) frequency in tumor DNA, suggesting that perhaps it’s not a driver mutation.
Recurrence of Mutations in Other AMLs
The other measure of importance of a somatic mutation is recurrence in other tumors of the same type. Thus, we screened for the presence of validated somatic mutations in a panel of 187 additional leukemia patients to see if any were recurrent. Most, unfortunately, were not. However, two variants were found in other samples, suggesting an important role in the development of AML. One was a noncoding conserved mutation (tier 2) on chromosome 10 which was detected in one other sample. Recurrence in just one other sample might not seem impressive, but by our estimation, the odds of such an event happening by chance are 1.1 x 10^-9. Thus, we may have uncovered a noncoding functional mutation that contributes to carcinogenesis via an as-yet-unknown mechanism.
The other was a nonsynonymous (tier 1) mutation in IDH1 at residue 132. Sixteen of 187 other leukemia samples carried mutations at the same residue in IDH1, suggesting an important role for this gene in the development of AML. Somatic mutations in IDH1 were recently characterized in glioblastoma (GBM) by our friends at Johns Hopkins, but this is the first time that IDH1 mutations were described in AML.
Conclusions: Lots of Passengers, Not Many Drivers
After sequencing the complete tumor genomes of two AML patients, we have estimated that these cancers carry an estimated 750 somatic events. Most such events will be background passenger mutations, acquired in the progenitor tumor cell before it became cancerous. Admittedly, that means there’s much more work to do to fully characterize the sequence changes underlying development of AML and other cancers. Our group is eager for the challenge. With the ever-growing throughput of the Illumina platform and our automated pipelines for whole-cancer-genome analysis, we hope to sequence at least a hundred more cancers in the coming year.
References
Mardis, E., Ding, L., Dooling, D., Larson, D., McLellan, M., Chen, K., Koboldt, D., Fulton, R., Delehaunty, K., McGrath, S., Fulton, L., Locke, D., Magrini, V., Abbott, R., Vickery, T., Reed, J., Robinson, J., Wylie, T., Smith, S., Carmichael, L., Eldred, J., Harris, C., Walker, J., Peck, J., Du, F., Dukes, A., Sanderson, G., Brummett, A., Clark, E., McMichael, J., Meyer, R., Schindler, J., Pohl, C., Wallis, J., Shi, X., Lin, L., Schmidt, H., Tang, Y., Haipek, C., Wiechert, M., Ivy, J., Kalicki, J., Elliott, G., Ries, R., Payton, J., Westervelt, P., Tomasson, M., Watson, M., Baty, J., Heath, S., Shannon, W., Nagarajan, R., Link, D., Walter, M., Graubert, T., DiPersio, J., Wilson, R., & Ley, T. (2009). Recurring Mutations Found by Sequencing an Acute Myeloid Leukemia Genome New England Journal of Medicine DOI: 10.1056/NEJMoa0903840

Outstanding work. Just one question did you sequence another control tissue ? If so, what was the level of sequence identity between the two normal tissues ? Any surprizes?