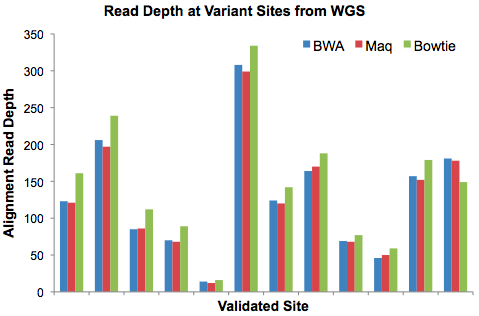
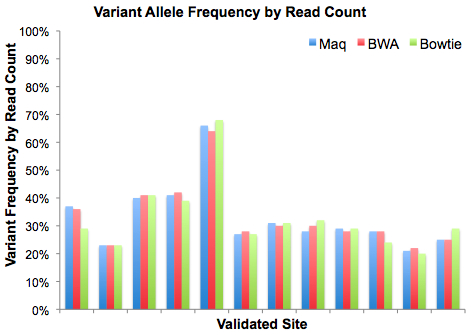
In recent weeks I’ve had conversations with many people in the NGS community who are attempting to call variants, accurately, in Illumina/Solexa data. Part of it stems from VarScan, my SNP and indel caller for next-gen sequencing data that works with Bowtie, Novoalign, cross_match, and other aligners.
Another part of it stems from my involvement in 1,000 Genomes Pilot 3, for which several participants have applied their own variant detection pipelines to the same dataset. Last month, Goncalo Abecasis, with input from David Craig, Heng Li, Gerton Lunter, and Fiona Hyland, proposed an exercise comparing several read mappers on real and simulated ABI SOLiD and Illumina/Solexa data. The initial list of aligners – Maq, BWA, Stampy, BFAST, BioScope, and KARMA – demonstrated just how rapidly the field has grown since my aligner comparison last year at AGBT. I’d looked at Maq and BFAST, and knew about (but hadn’t tried) BWA, but the others on the list (Stampy, BioScope, and KARMA) were ones I’d never heard of.
I proposed adding three aligners to the list: Bowtie and Novoalign for Illumina data, and SHRiMP for SOLiD data. My suggestions were politely declined by Richard Durbin (WTSI), who said “In our hands Bowtie doesn’t seem accurate enough for variant calling. It is a great tool for fast assignment of reads for some other purposes. Novoalign is accurate and good, but perhaps a little slow. SHRiMP is also I think very slow.”
Personally, I think that Bowtie works very well for variant calling, I know of several groups who are using it for that exact purpose. And while Novoalign *is* a bit slow, in my experience it’s just as fast as Maq, one of the two aligners out of Durbin’s lab that were already on the list. Of course, Maq remains the most widely used tool for Illumina data (for now), and that’s an important consideration. Most NGS analysts know and love Maq as much as I do.
Balancing Speed and Sensitivity
However, these assessments bring into focus the key issue surrounding short read alignment for variant detection – finding the balance between speed and sensitivity. Bowtie and Novoalign exemplify this well. Bowtie is ultrafast – the fastest short read aligner I’ve used – and maps an entire lane (~15m reads) in just 1-2 hours. Yet in my experience, it places slightly fewer reads than BWA/Maq. And it performs only ungapped alignments, so indels won’t be detected. In contrast, Novoalign typically maps more reads than Maq and BWA, seems very accurate, and remains one of the few aligners to allow gaps in fragment-end reads. In general, my comparisons demonstrated that Novoalign speed is comparable to Maq on typical datasets. However, longer reads and lower-quality data can make Novoalign very slow indeed. The ultimate short read aligner, in my opinion, would have Bowtie-like speed, Novoalign-like sensitivity, and the widespread community support that Maq enjoys.
Ask the Guru: Heng Li
Heng Li, who led development of both Maq and BWA, told me that he’s not worried about sensitivity. “Most aligners nowadays are sensitive enough,” Heng wrote to me in an e-mail this week. “For detecting variations, specificity is of more importance. Nonetheless, how much wrong alignments may contribute to wrong SNPs is an open question. As long as alignment errors are random, more wrong alignments may not necessarily lead to worse SNP calls.” Clearly, he has already given some thought to these issues. If we’re lucky, Heng Li may begin to address these open questions in his new post at the Broad Institute.
Underlying Causes of False Positives
Read mis-alignment would not be a serious problem if it occurred randomly across the genome. The trouble is that wrong alignments don’t seem to be random, at least in my experience. In projects like TCGA Ovarian, we see numerous false positives (particularly in tumors) that seem to arise from read mis-alignment. These typically manifest as clusters of variants, often present in each of a subset of reads whose true alignment is probably a paralogous region of the genome. It’s also possible that they’re caused by an indel, which (as Kiran Garimella of the Broad Institute recently showed) sometimes manifest as clusters of substitutions at several positions near one another. We can aggressively filter these by looking for clusters of predicted SNPs, but even better would be to remove the mis-alignments before variant calling even begins.
Read Mis-Alignment and Paired-End Sequencing
Here at WashU, we have a growing concern that the alignment scores for short reads are continually over-estimated. Often our manual reviewers find that reads supporting false-positives have mate pairs that align to a different chromosome altogether. In the absence of translocation events, when this occurs, one of the two reads is incorrectly placed, and any variant it supports is probably not real. Personally, I’d rather remove both reads in such situations, and rely on correctly mapped read pairs for detection of small variants.
The pervasive spread of paired-end sequencing is beginning to reveal just how often short aligners can get it wrong. The corollary here is that taking read pair information into account during alignment is of critical importance, and those hopeful short read aligners that don’t do it yet (crossmatch, for example) are destined for inferiority.
High-Throughput Sequencing: Speed Matters
Yet what I’m learning from discussions with others in the community – particularly the growing surge of users making the leap from Maq to BWA – is that speed matters. With Illumina machines cranking out 20 gigabases in a single run, and projects like the 1,000 Genomes generating terabytes of sequence over the course of months, we can’t afford to be using the slower aligners, no matter their sensitivity. At worst, we might apply a two-stage approach to alignment that rapidly maps reads that precisely match the reference, and passes only the variant reads to a more sensitive aligner for mapping.
Of course, as a colleague of mine recently joked, by the time we write the perfect aligner, Pac Bio will have come along and sequenced the entire genome, kilobases at a time.