Until recently, Maq has provided the central alignment/assembly/variant-detection functionality for our Illumina pipeline. As technologies and algorithms evolve, however, we continue to investigate possible improvements. Heng Li’s sequel to Maq, called BWA, utilizes the incredibly fast Burrows-Wheeler indexing algorithm to speed up alignment time by orders of magnitude. Also, BWA generates alignments in SAM/BAM format by default, which is convenient for our large-scale sequencing projects where BAM files are becoming the standard format.
These features, along with our impression that Heng Li and company do not plan future updates to Maq, lead me to infer that BWA is the heir-apparent for our Illumina pipeline. Before the transition, however, we must compare Maq results with BWA results on the same dataset, to identify any differences that may affect downstream analysis. Also, we are continuing to evaluate other aligners, especially Bowtie, which offer comparable or even better speed at short read alignment.
Test Data: WGS and Targeted Sequencing of a Single Sample
We have a sample in-house for which we performed whole genome sequencing (WGS) and subsequently validated numerous novel variants. We also performed capture-based targeted resequencing (Illumina 2x75bp PE) of 6,000 genes in the same sample. To compare the performance of BWA, Maq, and Bowtie, we aligned the capture data with each tool separately, and looked at about a dozen sites where we’d validated novel variants from WGS.
Sensitivity – Total Reads Mapped
Here’s a histogram of the read depth at each of the 12 variant sites by aligner:
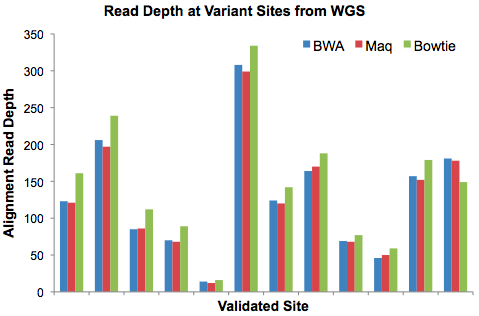
These results surprised me. Based on previous experience, I’d guessed that Maq would yield the highest depth, followed by BWA, and then Bowtie. Instead, with one exception, it was the other way around – Bowtie was more sensitive than BWA, which in turn was more sensitive than Maq. Yet these differences were relatively minor; overall, the coverage seems very comparable across all three aligners. I think that’s good news.
Variant Frequency by Read Count
Next, we looked at the observed variant frequencies, calculated as the relative fraction of reads supporting reference or variant alleles.
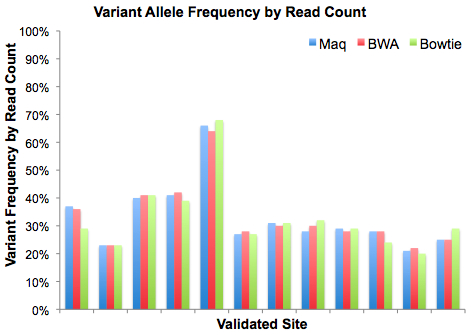
When it comes to variant frequency, Maq and BWA yield almost identical results (despite slight coverage disparities). Bowtie yielded slighly higher frequencies in some cases, slightly lower frequencies in others. Again, these were very minor differences from three very different alignment algorithms, suggesting that each of them yields fairly robust results.
Farewell to Maq
Unfortunately, the results of my analysis do not bode well for Maq, only because Maq took a few days to align data that BWA and Bowtie processed in a matter of hours. So which Burrows-Wheeler aligner will prevail? It’s difficult to say. As far as SNP detection goes, BWA and Bowtie seem comparable.
Which targeted capture platform/approach are you using? Any comments on how well it has (or has not) performed?
Any inkling of how robust these rankings would be to read length?
Dear Dan,
Nice post. You might find a poster we presented at CSHL on this topic informative too. All the best,
http://www.depristo.com/research/publications.php
Mark DePristo
Keith, thanks for the question. We’re using an oligo based solution-phase capture developed here that’s called WUCap. I can’t provide many details, as there’s a manuscript about to be submitted. However, Jon Armstrong gave a presentation on it at AGBT and was interviewed for an article in Genome Technology about capture methods.
Our Tech Development group is still refining the protocols, but thus far WUCap seems to perform comparably to Nimblegen capture, according to the data I’ve seen. Like most capture->NGS pipelines, we’ve encountered issues with repeats, missing probes, and GC bias, but overall I can tell you that the variant calling seems dramatically improved over PCR-based targeted sequencing.