As we continue to apply next-generation sequencing technologies to cancer genomes, we’re discovering hundreds of putative somatic mutations. Typically we run these through our annotation pipeline, which identifies variants affecting coding regions, splice sites, and evolutionarily conserved sequences. While a growing body of evidence suggests that much of the functional variation in humans lies outside these regions, inferring the functional impact of non-coding mutations remains challenging.
I just read excellent review by Lee, Yue, and Zhang in Human Genetics on Analytical methods for inferring functional effects of single base pair substitutions in human cancers. In it, the authors review bioinformatics approaches for evaluating single-base substitutions in both coding and noncoding regions, with an emphasis on large cancer resequencing efforts including the Tumor Sequencing Project, the Cancer Genome Atlas (TCGA), and the AML1 cancer genome. Several approaches to evaluating single-base mutations were presented:
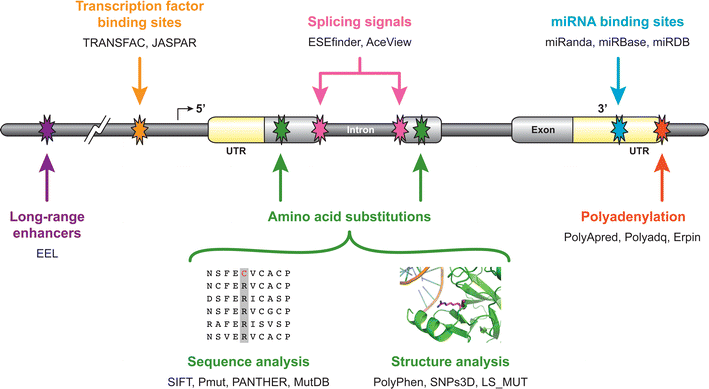
Credit: Lee W, Yue P, Zhang Z. Hum Genet. 2009
Coding Mutations: The Frequency Approach
The vast majority of somatic mutations in cancer are believed to be silent “passenger” mutations, which accumulate in cells but do not contribute to tumor growth and development. To find the smaller but more important “driver” mutations, several groups have applied a frequency approach. The idea is that mutations driving cancer are positively selected for during tumor development, and so their frequencies should be higher than those of passenger mutations. By assessing the incidence of a particular mutation or the frequency at which certain genes/pathways are altered across multiple patients (of the same tumor type), it should be possible to identify key mutations. Essentially, this approach uses recurrence as a metric for mutation significance.
Coding Mutations: The Amino Acid Approach
The frequency-based approach requires a significant number of samples, and likely favors driver mutations that are common across a patient population. Another way to go is bioinformatic analysis of individual protein-altering (nonsynonymous) mutations to score their probable effect on the protein. The widely used SIFT algorithm, for example, assesses the impact of NSS mutations using evolutionary conservation. Presumably, deleterious mutations are acted against by natural selection, and thus their amino acid residues should be conserved across species. Another algorithm, PolyPhen, applies protein structural information in a rule-based system to evaluate whether a mutation affects key protein domains. In a similar fashion, other tools assess whether mutations fall within annotated Pfam domains or signaling peptides of the encoded protein.
Coding Mutations: Classification Systems
Distinguishing driver from passenger mutations seems like a problem well suited for machine learning approaches. Indeed, groups have developed random-forest algorithms and support vector machine (SVM) approaches for this task as well. For any given mutation, a number of features (protein domain, conservation score, secondary structure, etc.) can be collected. Then, a positive training set (likely driver mutations from COSMIC) and a negative training set (common, presumably-neutral mutations) are used to build a classification system. Dave Larson in my group has applied a similar approach to distinguish between germline and somatic mutations.
Noncoding Mutations: Regulatory SNPs
Identifying functional SNPs outside of coding regions remains a challenge. One obvious category of such variants is “regulatory SNPs” whose alleles modulate the expression of nearby genes. Recently, groups have leveraged high-throughput gene expression and genotyping technologies to perform genome-wide assocation studies of gene expression, in which gene expression levels are considered a quantitative trait and significant correlations to SNP variation are identified. These studies have identified hundreds of cis-regulatory and trans-regulatory SNPs with moderate to large effects on gene expression. In cancer, some regulatory variants have already been identified. A SNP in the MDM2 gene (SNP309), for example, causes over-expression of this p53-suppressor and increases risk of colorectal cancer. Clearly, noncoding mutations affecting gene expression are of high interest in cancer genetics.
Noncoding Mutations: Post-Transcriptional Modifiers
Mutations that affect post-transcription processes – like splicing, polyadenylation, or the binding of regulatory RNAs – are also likely suspects in cancer development. Several mutations in BRCA1, for example, have been shown to alter that gene’s splicing and increase cancer risk. Mutations in the 3′ UTR of HMGA2 contribute to carcinogenesis by removing the binding sites for the let-7 microRNA, which normally represses this oncogene. To identify mutations in categories like these, we rely largely on computational tools that identify and score “motifs” – DNA sequences bound by micro-RNAs, splice enhancers, and other post-transcriptional players. Unfortunately, successes in this arena are few and far between. Nevertheless, I anticipate that interest in post-transcriptional modifiers will grow substantially as we learn more about alternative splicing, micro-RNA repression, and other relatively new areas of genetics.
The theme of the review article seemed to be this: a plethora of tools exist for evaluating mutations in cancer, and we’d better start leveraging them now. Whole-genome sequencing of cancer genomes is going to provide a flood of candidate somatic mutations. If only there were a single tool that applied all of the approaches above to help us prioritize them.
References:
Lee, W., Yue, P., & Zhang, Z. (2009). Analytical methods for inferring functional effects of single base pair substitutions in human cancers Human Genetics DOI: 10.1007/s00439-009-0677-y
Ding, L., Getz, G., Wheeler, D., Mardis, E., et al. (2008). Somatic mutations affect key pathways in lung adenocarcinoma Nature, 455 (7216), 1069-1075 DOI: 10.1038/nature07423
Ley, T., Mardis, E., Ding, L., et al. (2008). DNA sequencing of a cytogenetically normal acute myeloid leukaemia genome Nature, 456 (7218), 66-72 DOI: 10.1038/nature07485
The Cancer Genome Atlas Research Network. (2008). Comprehensive genomic characterization defines human glioblastoma genes and core pathways Nature, 455 (7216), 1061-1068 DOI: 10.1038/nature07385