The short read aligner Bowtie has gone legitimate, with a publication last month in Genome Biology and a mention by GT’s Daily Scan. While it has yet to supplant Maq as the de facto standard for Illumina/Solexa processing, Bowtie remains one of my favorite short read aligners. It was the first tool (to my knowledge) to implement Burrows-Wheeler Transform indexing, a method fast enough that it was soon adopted by the makers of SOAP and Maq. With last week’s paper, I finally got an idea of how BWT works:
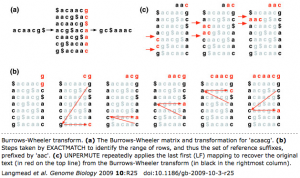 It was the first tool (to my knowledge) to implement Burrows-Wheeler Transform indexing, a method fast enough that it was soon adopted by the makers of SOAP and Maq. With last week’s paper, I finally got an idea of how BWT works.
It was the first tool (to my knowledge) to implement Burrows-Wheeler Transform indexing, a method fast enough that it was soon adopted by the makers of SOAP and Maq. With last week’s paper, I finally got an idea of how BWT works.
How Much Faster is Bowtie than Maq?
Based on my experience, I’d say it’s orders of magnitude faster, with only a slight hit in sensitivity. Here’s some data to back me up: CPU time to align a full flowcell (by lane) of 36-bp fragment-end reads to Hs36.
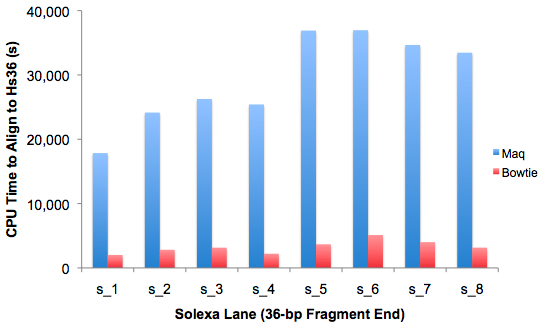
Figure 1. CPU time to align 1 flowcell of Illumina/Solexa 36-bp SE reads, by lane
On average, Maq took ~8 hours per lane to align the reads to Hs36, whereas Bowtie took just 54 minutes per lane.
New Paired-End Functionality
Also exciting is yesterday’s Bowtie update, which includes the much anticipated paired-end alignment mode. Paired-end mode is not only important for placing more reads, but also makes detection of structural variation with Bowtie all the more easier. While I haven’t yet evaluated this feature, if it’s done well, then Bowtie has become a serious player in the short read alignment game.