The short read aligner Bowtie has gone legitimate, with a publication last month in Genome Biology and a mention by GT’s Daily Scan. While it has yet to supplant Maq as the de facto standard for Illumina/Solexa processing, Bowtie remains one of my favorite short read aligners. It was the first tool (to my knowledge) to implement Burrows-Wheeler Transform indexing, a method fast enough that it was soon adopted by the makers of SOAP and Maq. With last week’s paper, I finally got an idea of how BWT works:
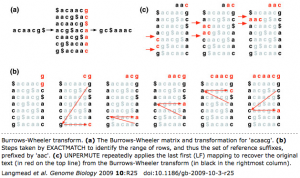 It was the first tool (to my knowledge) to implement Burrows-Wheeler Transform indexing, a method fast enough that it was soon adopted by the makers of SOAP and Maq. With last week’s paper, I finally got an idea of how BWT works.
It was the first tool (to my knowledge) to implement Burrows-Wheeler Transform indexing, a method fast enough that it was soon adopted by the makers of SOAP and Maq. With last week’s paper, I finally got an idea of how BWT works.
How Much Faster is Bowtie than Maq?
Based on my experience, I’d say it’s orders of magnitude faster, with only a slight hit in sensitivity. Here’s some data to back me up: CPU time to align a full flowcell (by lane) of 36-bp fragment-end reads to Hs36.
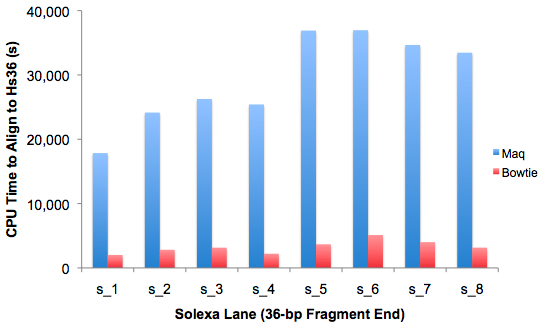
Figure 1. CPU time to align 1 flowcell of Illumina/Solexa 36-bp SE reads, by lane
On average, Maq took ~8 hours per lane to align the reads to Hs36, whereas Bowtie took just 54 minutes per lane.
New Paired-End Functionality
Also exciting is yesterday’s Bowtie update, which includes the much anticipated paired-end alignment mode. Paired-end mode is not only important for placing more reads, but also makes detection of structural variation with Bowtie all the more easier. While I haven’t yet evaluated this feature, if it’s done well, then Bowtie has become a serious player in the short read alignment game.
I know SOAP2 could generate solid results back to June last year. The first BWA was released also in June last year. Bowtie was released in August according to the SourceForge site. The key person behind SOAP2 is Tak Wah Lam, the first author of BWT-SW. If you have a look at his publication list in DBLP, you will find he has been in this field for years. I do not know when Bowtie is developed, but to the best of my knowledge, the three BWT-based aligners were largely developed in parallel. As BWA learns a lot from BWT-SW according to its manual page, I would more like to believe SOAP2 is the first BWT-based aligner.
Nice post, Dan! To “MB” – you write that you “would like to believe” that SOAP2 is the first BWT-based aligner, but publication is widely used in the scientific world to establish precedence. Otherwise anyone could claim his/her tool existed first. Ross Lippert was actually using the BWT for whole-genome alignment in 2005, and others used it previously too. But for short-read alignment, Bowtie is the first BWT algorithm to appear in the literature.
To Steven: I really respect your significant contribution to the field and I agree that publication is used to estabilish the precedence. However, publication is just a record of history and can be inaccurate. I am more interested in the true history. Ruiqiang Li, the SOAP developer, sent slides about SOAP2 to the 1000genomes mailing list on May 27, 2008. From the slides we know SOAP2 is working, even for paired-end reads and gapped alignment which is still missing in Bowtie. Back to that time, Bowtie and BWA were unknown to us, at least to me. Maybe Ruiqiang is a perfectionist to only publish perfect package, maybe he was too busy to write up a paper, maybe the collaborators wanted to keep it secret (as it is not open source till today), or maybe SOAP2 was not fully ready at that time, but I think it is anyway appropriate to acknowledge his contribution, the contribution from a large country but frequently ignored by the rest of the world.
I would seriously apologize if someone pointed me to the announcement/release of either Bowtie or BWA a month before May 27, 2008.