Exome sequencing has proven to be an incredibly powerful tool in the field of cancer genomics. It’s not just for mutations, either. The extensive coverage and continued improvement of current exome kits enables them to survey >90% of coding regions, as well as microRNAs, untranslated regions (UTRs), and other parts of the genome likely to harbor functional variation. Enrichment for such regions, coupled with the growing throughput of next-gen sequencing, yields sufficient coverage to identify small sequence variants (SNVs and indels) with great accuracy. New analysis methods tailored to exome data have also shown that it’s possible to search for gene fusions and even copy number alterations (CNAs).
One of our analysis tools, VarScan 2, offers a comprehensive suite of analysis tools for cancer exome sequencing. Given exome data for a tumor and its matched (usually blood) normal sample, VarScan 2 identifies SNVs and indels and classifies them according to somatic status: Germline (inherited), loss of heterozygosity (LOH), and somatic (acquired). Also, by directly comparing exome sequence depth, VarScan 2 detects changes in copy number in the tumor relative to the normal. This works well because the two samples (tumor and normal) are largely identical at a genetic level, and usually they’re sequenced under identical protocols. We published the algorithm in Genome Research late last year.
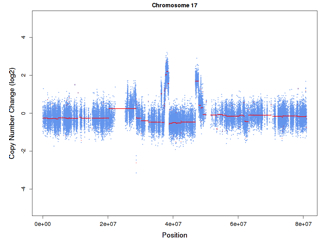
The output of the algorithm is a set of regions, each with chromosomal coordinates and a log2-ratio of copy number change. It’s very similar to array-based copy number data, and amenable to the same segmentation methods. Thus, we apply circular binary segmentation (CBS) to smooth the data and detect significant copy number change-points.
See the example at right, which is from a Her2+ breast cancer. You’re looking at several alterations on chromosome 17; the right-most peak is a focal amplification of ERBB2, the gene encoding the Her2 receptor.
Integrative Analysis of Tumor Exomes
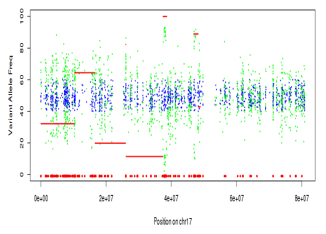
Cancer genomes often harbor numerous types of genetic alterations – mutations, structural variation, gene conversion events, etc. No single approach can survey everything at once, but exome sequencing is advantageous because mutations, copy number changes, and zygosity changes can be characterized simultaneously. Look at the allele frequency plot for the same chromosome offers some clarification:
- A large hemizygous deletion with concurrent LOH (red lines) spanning the first ~20 Mbp of the chromosome including the TP53 gene.
- Allele frequency shifts under the amplifications: it looks like ERBB2 has 2 extra copies, and the other amplification has 3 or 4.
- Another possible event spanning the last ~25 Mbp of the chromosome, possibly a deletion with incomplete LOH.
This type of analysis, integrated with the somatic mutation data, helps distinguish driver genes that are targeted for alteration in tumors, e.g. amplification of ERBB2 and deletion with LOH of TP53.
Why Survey Exome Copy Number?
I can think of at least three reasons why exome-based copy number analysis is appealing:
- High resolution. Surprisingly, between the number of gene targets and the “shoulder effect” of hybrid capture, exome data provides fairly comprehensive coverage genome-wide. Thus, exome-based CNA analysis provides a high-resolution survey of copy number alterations in a tumor genome.
- Unexploited datasets. In my post cancer genome and exome sequencing in 2011, I summarized dozens of studies of leukemia, melanoma, carcinoma, and other cancers enabled by exome sequencing. None of these, to my knowledge, included a copy number analysis from exome data. Most did array-based CNA analysis (usually SNP arrays), which provides more uniform coverage of the genome, but has far lower resolution than exome-based CNA analysis.
- Lots of data will be out there. The exomes of thousands of tumors have been sequenced already, and many of these datasets are in the public domain. The Cancer Genome Atlas Consortium, for example, has sequenced >1,000 tumor-normal exome pairs. Its study of ovarian carcinoma was published last year; studies of breast carcinoma, endometrial cancer, acute leukemia, and other tumor types will likely be out within a year.
Just a friendly reminder for everyone mining these rich datasets: the data are made available with the understanding that you can’t publish on them before the “marker paper” from TCGA comes out for that dataset. But you can certainly get started.
References
Koboldt DC, Zhang Q, Larson DE, Shen D, McLellan MD, Lin L, Miller CA, Mardis ER, Ding L, & Wilson RK (2012). VarScan 2: Somatic mutation and copy number alteration discovery in cancer by exome sequencing. Genome research PMID: 22300766