Contents: Whole-genome Sequencing • Recurrently Mutations • Clonal Evolution • References
Myelodysplastic syndromes (MDS) are a group of disorders of ineffective blood production and the most common cause of acquired bone marrow failure in adults. One-third of cases go on to develop secondary AML (sAML), yet there remains uncertainty among patients, insurers, and funding agencies about whether the myelodysplastic syndromes are actually cancers. A study online today at the New England Journal of Medicine has characterized the genetic evolution from MDS to sAML using whole-genome sequencing.
Whole-genome Sequencing of sAML
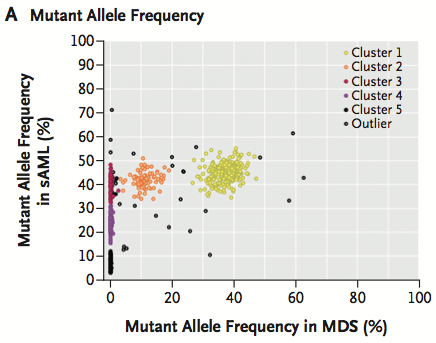
Matthew J. Walter and colleagues of the Washington University School of Medicine performed whole-genome sequencing of tumor samples and matched normal DNA from seven patients with secondary AML. For each subject, hundreds of somatic mutations were genotyped in sAML and MDS-stage samples to characterize the clonal architecture of each tumor. Figure 1A from the paper demonstrates the resolution that can be obtained from deep resequencing of somatic mutations in both sAML and MDS samples:
Notice the five clusters (differently colored) representing five clonal populations. In yellow (cluster 1) are mutations present in virtually all cells of both the MDS and the sAML sample. In orange (cluster 2) are mutations present at low frequency in MDS but enriched in sAML. Three more clusters (red, purple, and black) along the y-axis represent mutations that were absent in the MDS sample but acquired during the progression to sAML. The patterns of these mutations suggest that sAML evolved from a clonal population of MDS cells that acquired new mutations along the way.
Identification of Recurrently Mutated Genes
In the very near future, it may become feasible and cost-effective to perform whole-genome sequencing (WGS) on hundreds or thousands of tumors of a certain type to exhaustively identify recurrently mutated genes. Until then, WGS of a discovery cohort followed by extension screening in a larger cohort offers a powerful and cost-effective strategy. Two genes were already recurrently mutated in the 7 WGS cases: RUNX1, a known myeloid tumor suppressor, and UMODL1, for which mutations were recently reported in multiple myeloma and ovarian cancer. The authors extended their findings via targeted screening for additional coding mutations in 200 AML cases. This enabled the identification of 9 more recurrently mutated genes, for a total of 11.
Recurrently Mutated Genes in MDS and sAML
| Gene | Mutation(s) |
| CDH23 | 1235insL |
| NPM1 | W288fs |
| PTPN11 | G60R |
| RUNX1 | G170fs; del21q22.11 |
| SMC3 | e8-1 splice |
| STAG2 | H738fs |
| TP53 | V272M |
| U2AF1 | S34F |
| UMODL1 | T533P; V882M |
| WT1 | D436E |
| ZSWIM4 | P18A |
Notably, four of the genes (CDH23, SMC3, UMODL1, and ZSWIM4) had not been implicated in MDS or AML. A specific codon (34) in U2AF1 harbored missense mutations in multiple AML tumors, suggesting a gain-of-function for the splicing factor encoded by that gene. The recurrent mutations in STAG2, a gene located on the X-chromosome, were all protein truncation mutations (nonsense or frameshift) suggesting that a loss-of-function of this gene contributes to MDS and AML pathogenesis.
Clonal Evolution: from MDS to AML
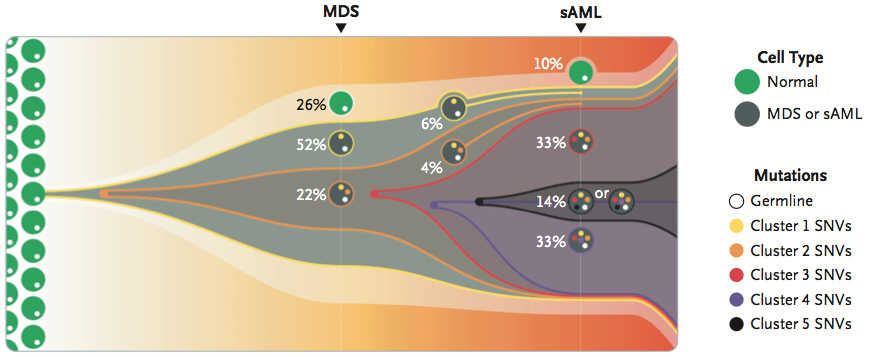
By characterizing mutations from secondary AML tumors in the MDS precursors for the same patient, the authors reconstructed the clonal architecture of the disease from early to advanced stages. The findings are summarized in Figure 2A:
In all 7 cases, the results suggest a linear model of clonal evolution, in which progression from MDS to sAML was characterized by persistence of a single founder clone (defined by ~200-700 mutations) and the outgrowth of at least one new subclone which contained dozens or hundreds of additional mutations. In other words, a single population of MDS cells underwent multiple rounds of mutation and selection, giving rise to multiple subpopulations present in full-blow secondary AML.
Please go read this fascinating study at the New England Journal of Medicine.
References
Walter MJ, Shen D, Ding L, Shao J, Koboldt DC, Chen K, Larson DE, McLellan MD, Dooling D, Abbott R, Fulton R, Magrini V, Schmidt H, Kalicki-Veizer J, O’Laughlin M, Fan X, Grillot M, Witowski S, Heath S, Frater JL, Eades W, Tomasson M, Westervelt P, DiPersio JF, Link DC, Mardis ER, Ley TJ, Wilson RK, & Graubert TA (2012). Clonal architecture of secondary acute myeloid leukemia New England Journal of Medicine