Exome and whole-genome sequencing offer powerful assays for disease diagnosis in clinical settings. In theory, they can help uncover the de novo mutations or inherited alleles responsible for rare genetic diseases. Over the past few years, several groups have developed strategies for filtering or prioritizing variants based on their likelihood to cause disease. Our tool, MendelScan, scores variants based upon:
- Segregation with the phenotype under the expected inheritance model (dominant or recessive)
- Predicted impact on gene or regulatory elements
- Frequency of alleles in human populations
- Expression of the affected gene in tissue(s) of interest
Other tools such as pVAAST and KGGSeq take a similar approach. The problem is that most cases don’t come into the clinic with large, well-phenotyped family pedigrees. You might have a family quad (parents + sibling), a trio (parents), or even just the proband. As good as they are, variant prioritization tools will still return hundreds of candidate variants. Curating those in the context of the disease phenotypes can be a tedious and time-consuming process, which is one of the reasons why exome sequencing for Mendelian disorders can fail.
Leveraging Gene, Phenotype, and Disease Knowledge
A paper in AJHG from Mark Yandell’s group — the team that developed the VAAST — aims to combine the outputs of variant prioritization tools with information that we have in gene/disease/phenotype ontologies.
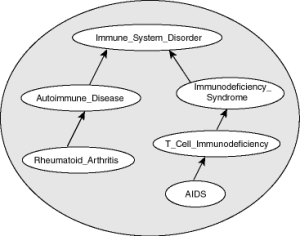
A basic (disease) ontology that reveals how AIDS is connected, in a manner of speaking, to rheumatoid arthritis
Ontologies are essentially knowledge databases in which the information (like the association of genes with pathways and processes) is represented as a graph. There are nodes (points) representing genes, phenotypes, or diseases, and then edges (connecting lines) between those nodes indicating that there’s evidence of a relationship. Their tool, called Phevor, leverages a few different ontologies:
- The Human Phenotype Ontology (HPO), which connects phenotypic abnormalities (e.g. diseases) to molecular and cellular processes
- The Mammalian Phenotype Browser (MPO), a similar resource but driven by animal models (it’s from Jackson Labs)
- The Disease Ontology (DO), which describes clinical characteristics of disease in a standard vocabulary
- The Gene Ontology (GO), which describes gene product attributes across species and categories
How the Ontologies Are Used
This ontology stuff has a steep learning curve, and I’m no expert, but let me give you an overview of what Phevor does. You provide it with some disease phenotype information, such as “hypothyroidism” (an example the authors provide), or an OMIM term. The first thing Phevor does is attempt to build a gene list using the HPO. Obviously it begins by looking for genes associated with that phenotype. If there are none, it looks at phenotypes connected to yours, and ones connected to those, until it finds associated genes. It compiles all of the genes into a list.
These genes become the starting points for searching the other ontologies (MPO, DO, and GO). There’s some fancy “ontology traversal” stuff that goes on, but in essence, the closer to a starting gene, or the more often that the same gene comes up, the higher it scores. This is useful because you may not have diseases or conditions that have genes associated with them already. Even if you do have candidate genes, this approach provides a rational and systematic way to expand your candidate gene list.
Scoring Each Gene
For each gene identified in the steps above, Phevor calculates two scores:
- A disease association score, which combines the score from the ontology search with the variant rank you got from your variant-prioritization tool of choice
- A healthy association score, which “summarizes the weight of evidence that the gene is not involved with the individual’s illness.” I understand the concept, but it was not clear how they calculate this.
The final Phevor score is the log10 of the ratio of disease score to healthy score.
Evaluating Phevor and Comparing to Other Tools
When you develop a tool that’s somewhat novel, it can be difficult to make useful comparisons of its performance to other analysis approaches. Even so, the authors’ choices here were a bit odd: they compared Phevor to “genome-wide search tools” VAAST and ANNOVAR, and “conservation-based variant-prioritization tools” (SIFT and PhastCons). This is an odd comparison, because VAAST is the only one that (in my opinion) truly does variant prioritization. ANNOVAR is a general-use annotation tool, SIFT evaluates only nonsynonymous mutations, and PhastCons just measures evolutionary conservation (the results of which are dependent on what other species are used).
Of these four tools, only VAAST attempts to do what I’d call variant prioritization. It’s also a tool from the Yandell lab, so not the most neutral choice. A better one might have been Ingenuity’s VariantAnalysis, but that’s a commercial tool. Still, I think VAAST was useful in illustrating what Phevor will do given a set of prioritized variants.
The figures in the main text are not terribly exciting, so I’ll summarize the results. Basically, when the authors used VAAST in real cases with known disease-causing alleles, 99% of the correct genes were ranked in the top 100, but none were in the top 10 (average rank: 83). When you apply Phevor to the VAAST results, 100% of correct genes moved up to the top 10 (average rank 1.8).
Variant Prioritization Gets You Only So Far
I think the header of this section nicely describes what most of us have come to know about exome sequencing in rare diseases. In many research situations and most clinical settings, there won’t be enough affected and unaffected samples from a large enough pedigree to truly narrow down to a single gene. Instead, you’ll be left with tens or hundreds of candidate genes, any one of which might be causal, and you’ll try not to think about the possibility that an incorrect pedigree or an undetected mutation (e.g. in a noncoding region) has already doomed your results.
Approaches like this one, which attempt to leverage knowledge buried deep in hard-to-use databases like GO and HPO, should be useful in sifting the wheat from the chaff.
References
Singleton MV, Guthery SL, Voelkerding KV, Chen K, Kennedy B, Margraf RL, Durtschi J, Eilbeck K, Reese MG, Jorde LB, Huff CD, & Yandell M (2014). Phevor combines multiple biomedical ontologies for accurate identification of disease-causing alleles in single individuals and small nuclear families. American journal of human genetics, 94 (4), 599-610 PMID: 24702956
[…] Ascending from the valley of despair. What can you if your main area is family studies and recruitment of small families is your focus? How can you escape the difficulties that are inherent to genetic studies in small dominant families? By changing the ground rules! First, small families carry additional genetic information compared to singletons, and if family recruitment is efficient, it may almost reach the efficiency of singleton recruitment, adding the increase in genetic information virtually at no cost. Second, small families may show you the value of non-established candidates (see “Flaws”). There is an entire cosmos of genetic variants that were only found in individual families before. Even small families can add to this if they add the crucial additional family. For this purpose, it is important that data are accessible for other researchers. Third, the field is evolving. The theoretical number of 100-200 variants identified in a small multiplex family is probably an overestimate – in fact we might get better and better at guessing the causative gene based on our gain of knowledge about the existing variation in the human genome and the particular phenotype. […]