A new study in PLoS ONE resequenced candidate genes in a cohort of the “healthy oldest-old” – individuals aged 85 or older that are healthy and have never been diagnosed with cancer, cardiovascular disease, Alzheimer’s, pulmonary disease, or diabetes. The idea is that these robust old-timers harbor genetic variants that reduce susceptibility to, or even protect against, the prevalent age-related disorders that tend to shorten lifespans. Demographic data suggest that less than 36% of the population of western nations will live to see 85, and only a third of these (12% overall) will do while remaining in good health. Since longevity is highly heritable (~25%), it stands to reason that genetics play a key role.
Tortoise Still Winning the Race
Intriguingly, despite the sum of human technological achievements – in agriculture, sanitation, medicine, etc. – our maximum observed lifespan (122 years) is not the longest on the planet even among animals. Indeed, the authors point out that rougheye rockfish, bowhead whales, red sea urchins, and Galapagos tortoises easily outlive us, with lifespans of 150-200 years. We can extend the lifespan of other animals – mice, by putting them on reduced-calorie diets, and C. elegans, by inhibiting expression of insulin/IGF receptor daf-2 – but can’t seem to change our own.
Rounding Up the Usual Suspects (Genes)
Next, the authors selected 24 candidate genes known to be involved in age-related processes. These included genes implicated in dietary restriction (SIRT1/3, UCP2/3, PPARG), autophagy (FRAP1, BECN1), stem cell activation (NOTCH1, DLL1), progeria syndromes (LMNA, ZMPSTE24, KL), tumor suppression (TP53, ING1, CDKN2A), and DNA methylation (TRDMT1, DNMT3A/B). Also included were the human homologs of several genes known to be differentially expressed in long-lived daf-2 mutant worms: IGF1R (growth factor receptor), SCD and APOB (lipid metabolism), and CRYAB and HSPB2 (heat shock proteins). Such an esoteric gene list allowed the authors to screen for variants across a wide range of gene functions and biological pathways that might contribute to longevity.
Ye Olde Candidate Gene Resequencing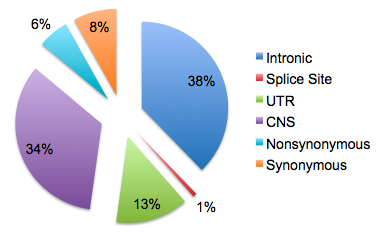
Some 716 PCR amplicons were designed to isolate the exons, 5′ and 3′ UTRs, 1.5 kbp promoters, intron-exon junctions, and selected conserved noncoding sequences (CNSs) for each of the 24 genes. Altogether some ~360 kbp of DNA was sequenced, bidirectionally, producing a grand total of ~35 million high quality (phred > 20) bases.
Variant detection with phred/phrap/polyphred and Mutation Surveyor identified 935 sequence variants (848 SNPs and 87 small indels), of which 59% were previously known to dbSNP. Unsurprisingly, the majority of variants found mapped to introns or conserved noncoding regions. About 50 novel coding SNPs were identified, though the authors point out that they were far less common (average MAF 1.6%) than the 80 or so previously known coding SNPs (average MAF 19%).
Tag SNPs: Leveraging the HapMap Resource
Here the authors took a rather puzzling turn and sought to compile a set of longevity tag SNPs by combining their data with the findings of the International HapMap Project. Only 12% of the combined variant set was shared between HapMap and the resequencing dataset, but that’s hardly surprising – HapMap variants were selected on the basis of high frequency (MAF > 5%), whereas many of the novel variants identified in this study were rare (in coding regions, MAF=1.6%). Thus the SNP sets are very likely to complement one another.
The authors selected 682 tag SNPs representing 1,550 non-redundant variants from the combined datasets (using LD > 0.8 for HapMap SNPs, LD >= 1 for resequencing SNPs). These were utilized to genotype a larger cohort (493 healthy oldest-old and 439 random controls), but unfortunately, the data was not shown. How disappointing! It seems to me that if the authors had found any significant association between their tag SNPs and longevity, that would have been an important result.
Common vs. Rare Variants: Is HapMap Enough?
One conclusion that was perhaps over-emphasized was that HapMap SNPs were inadequate to capture rare variation in the study population. Some 264 of the 935 variants identified by resequencing were singletons, i.e. present in just one individual, and only around 2.5% of these could be captured by HapMap tag SNPs using r-squared of 0.8. The authors conclude that “This shows that HapMap tagSNPs generally do not adequately represent, private re-sequencing SNPs. This analysis highlights a major challenge for genetic association studies. Using only HapMap SNPs, effects due to uncommon variants would often be missed.” Well, yes, but also, duh. HapMap was intended to represent common, and not rare, variation. Far more compelling would have been if the authors found rare variants actually associated with their phenotype of healthy aging. But alas…
The authors raise a fair point in that association studies cannot rely on the HapMap alone. To obtain the complete picture of genetic variation underlying a phenotype of interest requires a hybrid strategy that includes both common and rare variants. At some point this will require whole-genome resequencing of affected individuals, and for that, we’ll need something more than the 3730.
References
Halaschek-Wiener, J., Amirabbasi-Beik, M., Monfared, N., Pieczyk, M., Sailer, C., Kollar, A., Thomas, R., Agalaridis, G., Yamada, S., Oliveira, L., Collins, J., Meneilly, G., Marra, M., Madden, K., Le, N., Connors, J., & Brooks-Wilson, A. (2009). Genetic Variation in Healthy Oldest-Old PLoS ONE, 4 (8) DOI: 10.1371/journal.pone.0006641
Hi,
Nice post on a very interesting paper.
I selected your post as one of my “picks of the week” in molecular biology over at my blog (http://amontenegro.blogspot.com/2009/08/my-weekly-picks-from-researchblogging-2.html).
Cheers,
-A