 The past decade has been a turbulent one for the ream of genetic and genomic research. Since the completion of the human genome and beginning with the rise of next-generation sequencing, our community has seen some important changes in how we conduct research. As we look ahead to 2013 and beyond, we should remember for a moment what it took to get here.
The past decade has been a turbulent one for the ream of genetic and genomic research. Since the completion of the human genome and beginning with the rise of next-generation sequencing, our community has seen some important changes in how we conduct research. As we look ahead to 2013 and beyond, we should remember for a moment what it took to get here.
Part I: A New Hope
First came the advent of massively parallel sequencing, a technology arms race that began with two companies (Solexa and 454 Life Sciences) but went on to involve several others. These so-called “next generation” sequencing instruments enabled researchers to sequence and characterize genomes at an unprecedented pace. Resequencing the complete genome of an individual was suddenly within the realm of possibility. It still wasn’t cheap; I recall that our sequencing of AML1, the first cancer genome, required over 100 Solexa lanes.
Manufacturer’s note: Requires 20 informaticians and 200 CPUS (not included)
The rise of next-gen sequencing also spurred an era of intense bioinformatics development, as the research community scrambled to develop tools to handle the flood of short-read sequence data. A new crop of alignment and assembly algorithms was born. Variant calling was revamped and redefined. New viewers like IGV were needed to visualize the data.
Notably, due to the high instrument cost and staggering computational requirements, most next-gen sequencing took place at a handful of large-scale sequencing centers around the world. If you happened to work at one of these centers, these were golden days indeed. A single genome might merit a Nature paper, if you could ever finish analyzing it.
Big Sequencing Projects
The utility of this new sequencing power seemed endless. Imagine what we could discover with the complete genome sequences of many individuals. How many new species could be sequenced! Vast, ambitious efforts like the Cancer Genome Atlas, the 1,000 Genomes Project and the Human Gut Microbiome project were launched.
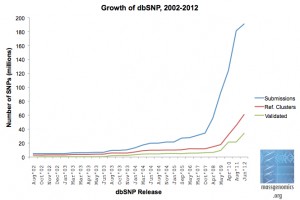
Of course we had to have some new databases to store all of this information, and compact yet flexible file formats (such as SAM/BAM and VCF) to store it in. The capabilities of next-gen sequencing would continue to surprise us: the Trace Archive couldn’t hold them, and even the Short Read Archive ultimately failed. Meanwhile, our knowledge of human genetic variation (as judged by the size of dbSNP) skyrocketed:
Part II: The Democratization of Sequencing
As sequencing technologies improved and new instruments came to market, the price of sequencing plummeted. More importantly, it became accessible: companies and core labs began to offer next-gen sequencing as a service. Instrument makers came out with lower-cost (and lower-throughput) benchtop sequencers that were much more affordable. Price wars ensued.
These developments brought about a second period of change in our field: the democratization of sequencing. The wide availability of high-throughput sequencing and ever-more robust and user-friendly analysis tools put the power of next-gen sequencing in the hands of the entire research community. Large-scale sequencing retained some advantages (capacity and experience) over the rest of the research community, but the gap was rapidly shrinking.
New NGS Applications
Also during this era, dozens of applications of NGS to study gene expression, DNA methylation, protein-DNA binding, and other biological processes were developed. Acronyms ending in -Seq were popping up everywhere. RNA-Seq, CHiP-Seq, MeDIP-Seq, you name it. The incredible throughout and low cost of sequencing made all kinds of new experiments possible. Many of these were used by the ENCODE project to characterize the content and function of the human genome.
Part III: Sequencing in the Clinic
Now I believe we’re entering yet another era, one that will ultimately have the most significant impact on human health: the transition of next-gen sequencing technology into the clinic. Already several leading cancer centers are using targeted capture and sequencing of tumors to guide the therapies of cancer patients. In the coming years, I expect to see broader adoption of DNA sequencing by clinicians. In the near future, the content of a patient’s genome will inform diagnosis, prognosis, and treatment. Personalized medicine, yeah baby!
Ethical Issues Abound
And while this is the logical next step for next-gen sequencing to take, it may also be the most complex and challenging time yet. And you thought developing and analyzing NGS data was the hard part! The social and moral and even financial hurdles to overcome in bringing genetic information into clinical settings are substantial. Ethical issues surrounding the ownership, predictive power, and privacy of patient genetic information abound. As highlighted in the final print issue of Genome Technology, investigators leading clinical sequencing efforts have faced ethical dilemmas already, such as the return of genetic information to family members in the event that a patient dies.
What If A Patient Dies?
In 2010, members of NHGRI’s ClinSeq study tried to follow up with a participant who’d enrolled two years earlier, and discovered that the patient had since passed away. At the time, they’d already sequenced and analyzed the patient’s exome as part of the study. The situation nicely illustrates some of the pivotal questions that researchers and clinicians will face when dealing with patients’ genetic information:
- Should the results of genetic studies be returned?
- What information should be returned?
- Who should be the recipient?
- How should the information be shared?
Another question is whether or not the family would want such information. In this case, the brother of the participant (and executor of estate) For help in addressing these questions, ClinSeq investigators consulted with NIH Clinical Center’s Bioethics Consultation Service. Eventually, the team reached the following conclusions:
- Certain results should be returned to the family, as they might have a bearing on the family’s health.
- Only variants with well-established clinical significance should be returned
- A single member of the deceased patient’s family (usually the executor of the estate) should receive the information in the family’s behalf
- The information should be shared in a face-to-face meeting with a genetic counselor.
Establishing Precedent for Bio-Ethics Decisions
While I’m not a bioethicist, I can see the logic behind these decisions, and I agree with them. No doubt each point was the center of vigorous debate among the ClinSeq team. I hate to say it, but theirs was a relatively easy situation, a softball. Genetic information was in hand and the family wanted it. Their decisions are helpful to the rest of us, because they begin to establish a precedent of how genetic information should be shared between patient and clinician.
References
Chan, B., Facio, F., Eidem, H., Hull, S., Biesecker, L., & Berkman, B. (2012). Genomic Inheritances: Disclosing Individual Research Results From Whole-Exome Sequencing to Deceased Participants’ Relatives The American Journal of Bioethics, 12 (10), 1-8 DOI: 10.1080/15265161.2012.699138
“The information should be shared in a face-to-face meeting with a genetic counselor.” — This sounds totally non-scalable. Once every newborn and cancer patient is sequenced…