Using genetic information to improve human health represents the central goal of biomedical research. Achieving it won’t be easy, but from a simplistic perspective, requires three steps.
- Cataloging the full extent of genetic variation in cells, tissues, and individuals. Efforts like the HapMap and 1,000 Genomes project are tackling this part, and judging by the explosive growth of databases like dbSNP and COSMIC, we’re well on our way.
- Correlating genetic variants to relevant phenotypes, such as disease predisposition and response to treatment. In principle, this is simply a matter of statistics: study a sufficient number of individuals, and the correlations should emerge.
- Understanding how (1) and (2) are related. In other words, how genetic and epigenetic changes exert their effects on a phenotype. This will undoubtedly be the hardest task of all.
An important step towards item #3 was made by the ENCODE consortium last month, when they published their guide to human genome content and function in the form of more than 30 research papers. Recently I highlighted some of the main ENCODE findings and their efforts to characterize regulatory variation in the human genome. Now, I’d like to summarize the landscape of transcription in human cells revealed by their efforts to characterize RNAs across 15 different human cell lines.
RNA represents the direct output of genetic information in a cell: the “message” of what’s encoded in its genome. As such, building a catalog of transcription in human cells is critical to understand how the genome functions. ENCODE sought to build a comprehensive dataset of RNA expression in these lines, interrogating both polyadenylated (poly-A) and non-poly-A RNA, long and short fragments, and even sequencing RNA isolated from specific cellular compartments (chromatin, nucleolus, and cytoplasm).
Pervasive Transcription Across the Genome
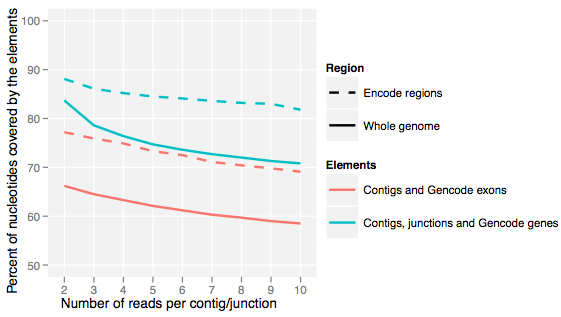
Perhaps the most surprising finding of the pilot project, and recently upheld by the main project, is that the human genome is pervasively transcribed: as many as 75% of bases are represented in at least one primary transcript:
Cumulatively, the authors observed that 74.7% of bases were covered by primary transcripts and 62.1% by processed transcripts. Interestingly, however, no single cell line expressed more than 57% of the union (complete set) of transcribed bases, suggesting that a signification portion of this transcription is cell-type specific.
Genes and Isoforms
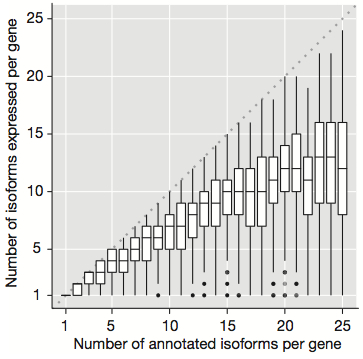
So, we’re all aware that there are more transcripts than genes: differences in transcription start sites and splicing, for example, have almost limitless potential for producing different isoforms of a given gene. Perhaps surprisingly, ENCODE found that genes are not expressed in a minimalist strategy. Look at the number of isoforms per gene:
By analyzing the different gene isoforms observed within and across cell lines, the authors also observed that:
- Alternative isoforms within a gene are not expressed at similar levels. Usually, one isoform predominates.
- About 3/4 of protein-coding genes expressed different dominant isoforms depending on the cell line and condition
- Across cell lines, differences in gene expression levels were more variable than differences in splicing.
Finally, the number of observed isoforms per gene correlated with the number of annotated isoforms (shown in the plot above), hitting a sort of plateau at 10-12 expressed isoforms per gene per cell line. That’s an incredible amount of variation, and it begins to suggest a theme brought home by this and many other studies: that even the basic elements of the genome (genes) are far more complex than we might have anticipated.
Characterization of Annotated and Novel Transcripts
One important element of the ENCODE project was its ambitious annotation effort (GENCODE) which sought to compile our current knowledge of splice junctions, transcripts, and genes. About 70% of these were detected by RNA-Seq across the 15 cell lines (85% of annotated exons were represented), and a large number of novel elements were present as well. These novel elements were concentrated around (but not limited to) gene regions, covering 78% of intronic bases (within genes) and 34% of intergenic bases (between genes). They add 20% more exons and 45% more transcripts to the GENCODE annotation.
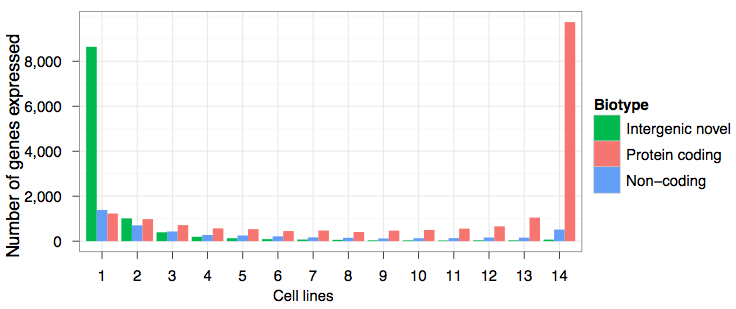
The distribution of gene expression was similar across cell lines, with protein-coding genes averaging higher expression levels than long non-coding RNAs (lncRNAs). Indeed, about 80% of observed lncRNAs were present at 1 fewer copies per cell, compared to 25% of protein-coding genes. The authors note that this may be a feature of lncRNA expression occurring only in a subset of cells, rather than a picture for all cells. In some cell lines, lncRNAs exhibited steady-state levels of expression comparable to those of protein-coding genes.
Also, lncRNAs were more likely to be specific to a single cell line (29%) than protein-coding genes 7%); in fact, more than half of protein-coding genes were constitutive, i.e. expressed in all cell lines.
Note that while most protein-coding genes were expressed in all cell lines, novel intergenic genes were predominantly expressed in a single cell line.
Alternative Initiation and Termination
The authors employed two experimental techniques to examine where transcription starts and stops in the genome:
- Cap-analysis of Gene Expression (CAGE), in which capped 5′ ends of RNAs were isolated and sequenced, and
- Paired-end Tag (PET) sequencing, in which short tags at the 5′ start and 3′ end of full mRNA transcripts are sequenced
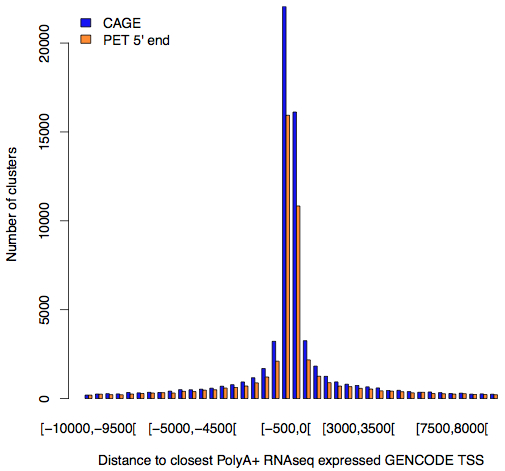
The transcription start sites (TSSs) identified by CAGE, and PET sequencing were compared to the list of 97,778 TSSs that were in the GENCODE annotation and expressed according to RNA-Seq:
About half of the CAGE-identified start sites were located within 500 bp of an annotated TSS that was expressed in RNA-Seq data. However, only 72% of all CAGE sequencing reads mapped near known TSSs, suggesting that the remaining 28% are from re-capping events or else represent a new class of transcription start site.
Characterization of Small Noncoding RNAs
Small noncoding RNAs (ncRNAs) have been an area of intense interest over the past several years. By sequencing short fragment size fractions in several cellular compartments, the ENCODE was able to characterize the expression and localization of several classes of small RNA:
- Micro-RNAs (miRNAs), which are short RNA fragments known to have a role in post-transcription regulation
- Transfer RNAs (tRNAs), the adapter molecules between mRNA and amino acids.
- Small nuclear RNAs (snRNAs) associated with the spliceosome
- Small nucleolar RNAs (snoRNAs), which guide chemical modifications (methylation and pseudouridylation) of ribosomal and transfer RNAs as well as snRNAs.
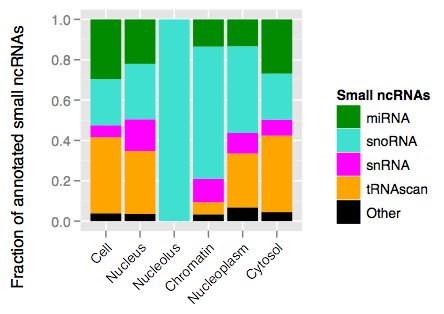
Given the function of these classes of RNAs, we might already have a guess as to the cellular compartments in which to find them. In the K562 cell line, here are the ncRNA compositions by cellular compartment:
Indeed, small RNAs were enriched in compartments where they perform their functions: miRNAs and tRNAs in the cytosol and snoRNAs in the nucleus. The abundance of snRNAs in chromatin-associated RNA suggests that, as has already been suggested, splicing predominantly occurs as a gene is being transcribed.
RNA Editing: Widespread or Rare?
RNA editing is a rare but remarkable phenomenon in which a base in an RNA transcript is changed after transcription. It’s also the subject of recent controversy. A 2011 paper in Science by Li et al had reported that RNA editing was widespread in the human transcriptome. At AGBT 2012, I attended a talk in which Vivian Cheung (the senior author) presented the work. To me, and some members of the audience, the flaws were kind of obvious. They hadn’t controlled for false positives in RNA-Seq data. As a result, they reported way too many events and across the full spectrum of possible substitutions. RNA editing is actually quite rare, and it’s almost always an A to G substitution.
Unsurprisingly, about a month later Science published comments from three independent groups challenging the findings. Kleinman and Majewski (McGill), for example, wrote “Li et al. did not properly control for a number of technical limitations of HTS and the downstream analysis of the data, resulting in an unacceptably high false-positive rate within this study.” It’s easy to understand how this could happen: variant calling with next-gen sequencing can be difficult, especially in RNA-Seq data because errors are more prevalent (due to reverse transcriptase errors and the challenges of making accurate gapped alignments of the reads).
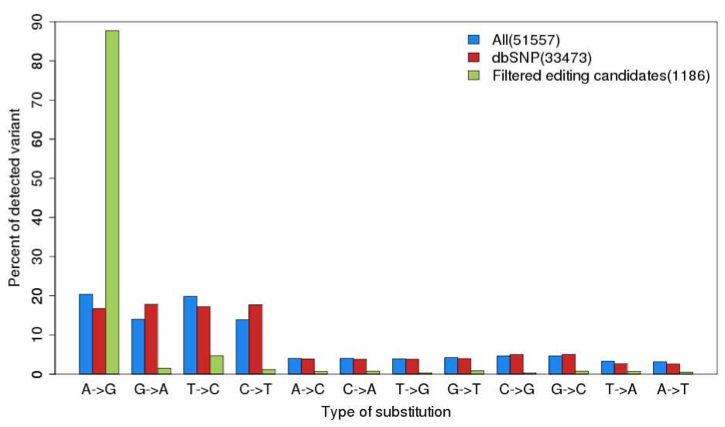
In this study, the ENCODE developed a pipeline to identify single nucleotide variants (SNVs) and remove sequencing artifacts. In the GM12878 cell line, which has been deeply resequenced, they called 51,557 “RNA consistent” SNVs of which 65% overlapped dbSNP positions (this indicates they’re germline variants, not RNA editing events). After removing those and applying even more stringent filters, the authors found 1,186 events in 430 genes, of which the vast majority (88%) were A to G changes.
In the words of ENCODE authors, “These results do not support a recent report of a substantial number of non-canonical SNV edits in the RNA of human lymphoblastoid cells.”
Conclusion: Relevance of the Transcriptome
In genetics and genomics, we often think of genetic variation at the level of the genome: inherited variants and acquired changes that may contribute to a phenotype such as disease susceptibility or response to treatment. Judging by the explosive growth of databases like dbSNP, DGV, and COSMIC over the past several years, we are rapidly on our way to cataloging the vast majority of genetic variation in the human genome. In other words, we have a pretty good idea of what the genetic differences are between one person and the next. We have a much less complete picture of what those variants do.
Understanding the “message layer” between a cell and its genome, through studies like these, will help us make the connection between genotype and phenotype.
References
Djebali S, Davis CA, Merkel A, Dobin A, Lassmann T, Mortazavi A, Tanzer A, Lagarde J, Lin W, Schlesinger F, Xue C, Marinov GK, Khatun J, Williams BA, Zaleski C, Rozowsky J, Röder M, Kokocinski F, Abdelhamid RF, Alioto T, Antoshechkin I, Baer MT, Bar NS, Batut P, Bell K, Bell I, Chakrabortty S, Chen X, Chrast J, Curado J, Derrien T, Drenkow J, Dumais E, Dumais J, Duttagupta R, Falconnet E, Fastuca M, Fejes-Toth K, Ferreira P, Foissac S, Fullwood MJ, Gao H, Gonzalez D, Gordon A, Gunawardena H, Howald C, Jha S, Johnson R, Kapranov P, King B, Kingswood C, Luo OJ, Park E, Persaud K, Preall JB, Ribeca P, Risk B, Robyr D, Sammeth M, Schaffer L, See LH, Shahab A, Skancke J, Suzuki AM, Takahashi H, Tilgner H, Trout D, Walters N, Wang H, Wrobel J, Yu Y, Ruan X, Hayashizaki Y, Harrow J, Gerstein M, Hubbard T, Reymond A, Antonarakis SE, Hannon G, Giddings MC, Ruan Y, Wold B, Carninci P, Guigó R, & Gingeras TR (2012). Landscape of transcription in human cells. Nature, 489 (7414), 101-8 PMID: 22955620
Just came across this post. I would point out one major issue with the interpretation of the ENCODE data that seems to have been missed in all of the hype. But I’ll preface it by saying I think the ENCODE generated data is incredibly useful and interesting. We should keep in mind that this is equivalent to a large scale screen and everything discovered is potentially biologically meaningful and worthy of follow-up, but they haven’t really shown anywhere near the 80% functional level that was stated in the papers and press releases. In fact their former lead data analyst confirmed in a blog post Q&A that really the figure is “somewhere between 20-80%”, with 80% representing an upper bound.
If you read the RNA-Seq papers though, I’m even a little leery of the 20% lower-bound. There is a huge amount of skew towards novel transcripts that are near known genes. If you look at the plots of data to get expression level, you’ll notice as well that there is substantial skew towards these novel transcripts being observed, on average, in less abundances that are less than a single transcript per cell…
RNA-Seq data is very sensitive, especially when cell lines are sequenced to incredible depths. We know that transcription is stochastically messy. Bindings sites for assembling the transcriptional machinery exist in locations that aren’t “real” bindings sites because evolutionarily it makes sense. There will always be a certain amount of noise, random transcription of non-functional RNA at very low levels, and this will tend to be clustered near to known genes. The novel transcripts need to be taken with a grain of salt. It would have been interesting to show the data at different filter cut-offs for expression level, and not just whether those transcripts were consistent across cell lines.