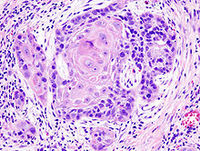
More than half a million new cases of head and neck squamous cell carcinoma (HNSCC) will occur in 2011, making it the 6th most common malignancy in the world. Two studies online at the journal Science survey the mutational landscape of this deadly cancer, which has a mortality rate of ~50%. They report frequent mutation of the NOTCH1 gene in HNSCC (11-15% of cases), and the patterns of these mutations suggest a tumor suppressive role. This observation is in stark contrast with many solid tumors and hematopoietic malignancies where Notch signaling is thought to play an oncogenic role. Moreover, it carries worrisome implications for Notch1 inhibitors, some of which have recently entered clinical trials.
Around 50,000 cases of HNSCC are diagnosed each year in the United States. The genomes of HNSCC bear many chromosomal aberrations, including amplifications targeting the CCND1 gene on chr11q13, and the epidermal growth factor receptor gene (EGFR) on chr7p11. Many of these tumors also exhibit genetic or epigenetic alterations of TP53 and CDKN2A, two well-known tumor suppressor genes. Tobacco and alcohol exposure are risk factors. More recently, HPV infection has emerged as a risk factor as well. Patients with HPV-associated tumors have a better chance of survival, indicating distinct biological features for this form of the disease.
Exome Sequencing of HNSCC
Stransky et al performed solution-phase capture and Illumina sequencing of 74 tumor-normal pairs, achieving 150-fold average depth of target regions and covering 87% of bases with at least 20 reads. They also performed SNP array-based copy number analyses. Common CCND1 amplifications, CDKN2A deletions, and rarer amplifications of MYC, EGFR, ERBB2, and CCNE1 suggested that their tumor set was genetically representative of HNSCC. Using exome data, the authors predicted ~130 coding mutations per tumor, of which 25% were synonymous changes.
Agrawal et al examined 32 tumor-normal exome pairs: 17 by Illumina sequencing, and 15 by SOLiD sequencing. They achieved 77-fold (Illumina) and 44-fold (SOLiD) average depth of target regions, with 90-92.6% of target bases covered by at least 10 reads. Most of the tumors (30 of 32) came from pre-treatment patients, and all were selected for >60% tumor cellularity. This latter selection was an important one, as the Stransky et al study had sequenced but not reported 18 additional tumor-normal pairs due to extensive stromal admixture.
HPV, TP53, and Tobacco Exposure
Taken together, both studies found that HPV-associated HNSCC truly represents a distinct disease at the molecular level, with a lower mutation rate (2.28 per megabase compared to 4.83 per megabase) than HPV-negative tumors. Further, none of the HPV-associated tumors carried TP53 mutations, whereas the gene was mutated in 62-78% of HPV-negative tumors. Only 18% of HPV-negative tumors had mutations in bona fide oncogenes, which is not good news for the prospect of targeted therapies. Mutation rates in smokers were higher than those of non-smokers. While Agrawal et al reported no evidence of tobacco exposure in their study, this might have been due to the limited sample size (32 tumors), because Stransky et al (with 74 tumors) observed an excess of G to T transversions at non-CpG islands, consistent with carcionogen-induced mutations.
Inactivating Mutations in Notch
Both studies made an interesting observation: an excess of mutations in the NOTCH1 gene, many of which were protein-truncating alterations. Further, several tumors had lost both copies of NOTCH1, either by mutation or large-scale deletion. These observations suggest that NOTCH1 is being inactivated in HNSCC. In contrast, the Notch signaling pathway is up-regulated in numerous human cancers, particularly hematopoietic tumors. The newly-established tumor suppressive role of NOTCH has important implications for cancer therapies, as several NOTCH pathway inhibitors have entered clinical trials. One of these trials was recently halted, partly because of treatment-associated skin cancers.
A number of other genes related to squamous cell differentiation proved to be mutated at significant frequencies, including NOTCH2, IRF6, TP64, RIPK4, CDH1, EZH2, Dicer1, and MLL2. Other mutations affected genes involved in calcium-sensing (RIMS2 and PLCO) or apoptosis (CASP8, DDX3X). These gene sets, and the NOTCH-related genes in particular, suggest an important role for normal squamous cell developmental pathways in the formation of squamous cell carcinoma.
References
Stransky N, Egloff AM, Tward AD, Kostic AD, Cibulskis K, Sivachenko A, Kryukov GV, Lawrence M, Sougnez C, McKenna A, Shefler E, Ramos AH, Stojanov P, Carter SL, Voet D, Cortés ML, Auclair D, Berger MF, Saksena G, Guiducci C, Onofrio R, Parkin M, Romkes M, Weissfeld JL, Seethala RR, Wang L, Rangel-Escareño C, Fernandez-Lopez JC, Hidalgo-Miranda A, Melendez-Zajgla J, Winckler W, Ardlie K, Gabriel SB, Meyerson M, Lander ES, Getz G, Golub TR, Garraway LA, & Grandis JR (2011). The Mutational Landscape of Head and Neck Squamous Cell Carcinoma. Science (New York, N.Y.) PMID: 21798893
Agrawal N, Frederick MJ, Pickering CR, Bettegowda C, Chang K, Li RJ, Fakhry C, Xie TX, Zhang J, Wang J, Zhang N, El-Naggar AK, Jasser SA, Weinstein JN, Treviño L, Drummond JA, Muzny DM, Wu Y, Wood LD, Hruban RH, Westra WH, Koch WM, Califano JA, Gibbs RA, Sidransky D, Vogelstein B, Velculescu VE, Papadopoulos N, Wheeler DA, Kinzler KW, & Myers JN (2011). Exome Sequencing of Head and Neck Squamous Cell Carcinoma Reveals Inactivating Mutations in NOTCH1. Science (New York, N.Y.) PMID: 21798897