The criminal justice system in the United States is rarely called an early adopter of new technology. Despite the major impact of forensic DNA testing over the last quarter-century, the tools deployed by most forensics laboratories are rudimentary by modern standards. If you compare what the FBI does with an unknown DNA sample to what 23andMe can do (faster and with lower costs), the differences are quite striking. The FBI can determine whether the DNA matches a reference sample (or an entry in its CODIS database) and that’s about it. In contrast, 23andMe can tell you how much Neanderthal is in your DNA and help you find your third cousins.
There’s encouraging news for the field of applied DNA forensics, as earlier this month the National Institute of Justice awarded an $825,000 grant to the Battelle Memorial Institute to “conduct feasibility and validation tests on a suite of new investigative tools that use next-generation sequencing.” It came to my attention thanks to Julia Karow’s feature article over at In Sequence.
Now seems like an excellent time to discuss the potential for NGS applications as well as some of the significant challenges.
How DNA is Currently Used in Forensics
Importantly, nearly all routine DNA testing for forensic purposes involves capillary electrophoresis of short tandem repeats, or STRs. Unlike SNPs, which tend to have two alleles, STRs have numerous alleles, defined by the number of repeats at each locus. Because they’re so polymorphic, STRs are well-suited for finding a “match” between two samples.
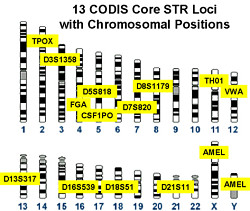
Theoretically speaking, about 10-12 carefully chosen STR loci are sufficient to identify an individual. In the U.S., the FBI uses a panel of 13 (plus the AMELX/AMELY loci for sex determination) for its COmbined DNA Indexing System (CODIS) database.
The current applications for DNA in forensics laboratories are focused on matters of the justice system:
- Matching a crime scene DNA sample to a reference (i.e. a cheek swab) to implicate a suspect, either directly or indirectly (i.e. using a close relative).
- Searching for a match among the profiles of criminals in federal databases like the FBI’s CODIS.
- Comparing samples from two crime scenes to learn if the same suspect was responsible
- Identification of human remains or found individuals in missing persons cases
Notably, although the STR profiles in CODIS are well-suited for matching DNA samples, they’re poorly suited for distinguishing an individual’s ancestry.
How DNA Could be Used in Forensics
DNA sequencing/genotyping technologies and applied human genetics have advanced considerably since the establishment of CODIS in the 1990’s. One obvious application, hinted at in the proposed plans of the Battelle/NIJ grant, is to empower the identification of “unknown” DNA samples (from crime scenes or missing persons cases) by building a profile of likely physical characteristics:
- Ancestry or continent of origin, based on ancestry-informative markers (AIMs). Back in 2006, when I worked in the lab of Raymond E. Miller, we developed a panel of about 25 SNPs that reliably distinguished between individuals of African, Asian, or European origins. With more SNPs one could trace ancestry with much higher resolution.
- Physical appearance, especially eye or hair color. These are not simple Mendelian traits, but with sufficient markers one could determine the probability of brown versus blonde hair, or blue versus brown eyes. Of course, it’s now possible to change your apparent hair or eye color using dyes and colored contacts, respectively, but let’s not go there.
- Deep familial matching is also possible, particularly with a sequencing-based assay that will detect rare variants unique to certain pedigrees.
- Faster, less expensive results may ultimately be what drives the adoption of new technologies. The turnaround time for DNA testing in most situations is slow, and the backlog is substantial.
Challenges of DNA Forensics
Given the obvious utility of new sequencing and genotyping technologies in forensic applications, you might be wondering: Why has it taken so long? Having done some work in this field, and discussed the matter with multiple law enforcement agencies, I can tell you some of the reasons.
Judicial Merit, Acceptance, and Precedence
First, adoption of new technologies is slow in the criminal justice system. To be useful in a criminal investigation, they must establish judicial precedence which takes a lot of vetting and a lot of time. No one wants to waste money on a technology that will be easily disputed by defense attorneys, right? The governmental agencies that operate most forensics laboratories exist as part of the justice system. They need to prove the merit of something before devoting substantial resources to it.
Established Infrastructure
It is difficult to emphasize how important the CODIS database is to forensic laboratories. Any technology that displaces capillary electrophoresis of STRs absolutely must provide CODIS profiles. Remember, CODIS has been around a long time (officially since the federal “DNA Identification Act of 1994”). As of last year, it contained 10.3 million offender profiles, 1.5 million arrestee profiles, and 493,500 forensic profiles. And it had assisted more than 200,000 investigations.
In short, CODIS is not going away. Next-gen sequencing is theoretically capable of generating STR profiles, but short read lengths make this more difficult.
Impure and Degraded Samples
In a research setting, we panic when we have sample contamination or low amounts of genomic DNA. We know it affects things like mapping rate and duplication rate of the resulting reads. Just a heads-up here: the DNA samples often encountered in forensic situations are a lab manager’s nightmare. They’re almost always mixtures of DNA from multiple people. They may be heavily degraded. They might not even be human in origin. Rarely will pure samples with good amounts of DNA come into a forensics laboratory.
This makes many of the desirable applications of forensic DNA sequencing a bit more difficult. Degraded DNA is typically fragmented and thus more difficult to sequence. Even when you can, the coverage might be incomplete. As I mentioned, the FBI uses 13 identification loci in its CODIS database. To add a profile, all 13 must be attempted, and at least 10 must be tested successfully.
Ethics and Privacy Concerns
You know how much I like to talk about the elephant in the room: ethical, legal, and social considerations for expanded DNA testing capabilities by government agencies. Groups such as the ACLU are already concerned with how DNA testing can provide “partial” (familial) matches. Imagine getting a knock at your door, and opening it to find two police detectives.
“Are you Mr. Smith?” one of them asks.
“Uh, yes,” you answer, already nervous from the sight of their badges and guns.
“We’re investigating a homicide,” says the other detective. “DNA evidence from the scene was a partial match to you. We’d like to ask you about all of your second and third cousins.”
This is a slight exaggeration, but with the power of next-gen sequencing and human genetics, it’s still plausible. We can’t live our live distrusting every single person and organization of authority, but we do need to have open conversations about implications of advanced DNA testing.
I tip my hat to the Battelle Institute for tackling both the hard science and the sticky issues. They have a bumpy road ahead.

Tapping into the power of NGS by forensic professionals is quite the move.
However, there is still work to be done on improving the traditional sample
collection tools being currently used in the forensics world, either for
reference and the crime scene samples. I strongly believe, that parallel path
is needed to have an improvement over what is being used currently. This work is a great start and looking forward to the outcome on what is needed to get NGS into the forensics world.