Many large-scale genetic studies are conducted in broad/homogenous population cohorts, like the panels chosen for the 1,000 Genomes Project. Yet there are advantages to studying smaller and more isolated founder populations, as exhibited in a study in this month’s Nature. Moltke et al report a genetic association study in the Greenlandic population, a small and historically isolated population (57,000 people). Of particular interest: type II diabetes (T2D), the prevalence of which has skyrocketed over the past 25 years.
Granted, T2D is a complex disease with many contributing factors. From a genetics perspective, the small and historically isolated population offers some advantages. The limited genetic diversity means that linkage disequilibrium is extended, which boosts the power of genetic association studies. Further, deleterious variants can reach higher frequencies due to the founder effect (genetic drift in a small population across many generations). This phenomenon has already been documented in the Finnish population, whose unique population history has contributed to the increased prevalence of at least 40 genetic disorders.
Part I: The Genome-Wide Association Study
In the current study, the authors began with a GWAS in ~2,700 samples collected from 12 locations in Greenland. They did the genotyping on the Illumina Metabochip, a customized SNP array with 200K variants marking regions implicated in metabolic, cardiovascular, and anthropometric traits. Notice how the authors are stacking odds in their favor? A founder population, with an increased prevalence of T2D, and a custom genotyping array to study the relevant traits.
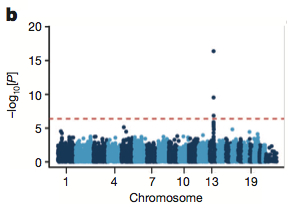
The authors employed a linear-mixed model to search for association of variant alleles with T2D status. You can see the hit on chromosome 13. That’s rs7330796, a variant in intron 11 of the TBC1D4 gene that was selected for inclusion on the Metabochip because it was in the top 5,000 candidate SNPs for association with waist-to-hip ratio.
Part II: Exome Sequencing and Fine Mapping
Even though this was a specialized SNP chip, it was unlikely that rs7330796 was the causal variant. It’s intronic, and not in a large LD block. So the authors performed exome sequencing on nine trios, and identified four coding variants in high LD with rs7330796. One of these was a nonsense variant, introducing an early stop codon (p.Arg684Ter) in the longer isoform of TBC1D4.
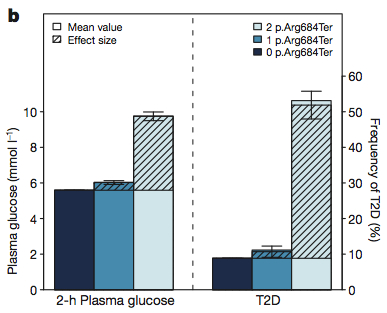
When genotyped in the main cohort, p.R684X was strongly associated with 2-h plasma glucose levels. When conditioning on rs733096 alleles, it was also associated with 2-h serum insulin levels.
Functional Assessments and Follow-up
The fascinating thing about this nonsense variant was that it had an almost Mendelian effect on plasma glucose levels. Heterozygous carriers had a slight elevation, but subjects homozygous for the alternate allele had significantly elevated plasma glucose. Strikingly, the frequency of T2D in homozygous carriers was also about 5x higher.
Thus, the p.R684X variant confers increased risk of a subset of diabetes that features deterioration of glucose homoeostasis. And with an odds ratio of 10.3, the variant exhibits an effect size much higher than any other variants associated with T2D to date.
Interestingly, though p.R684X has a MAF of ~17% in Greenlanders, the variant was essentially absent in other sequenced European populations (the 1,000 Genomes panels, 6500 ESP exomes, and 2,000 Danish exomes). Except for one Japanese sample from 1,000 Genomes. However, the Greenlandic population is an admixed population with European and Inuit heritage. Sure enough, when the authors looked at the Inuit population, they found the variant at a MAF of 23%. Thus, though this variant is not unique to Greenland, it’s far more common there than in other populations.
One criticism of classic GWAS approaches is that they typically identify genetic markers associated with certain phenotypes, not the causal variants themselves. This study went beyond that with exome sequencing and follow-up genotyping. The beauty is that TBC1D4 makes a lot of sense as a diabetes gene. It acts as a mediator of insulin-stimulated, Akt-induced glucose uptake by regulating the GLUT4 transporter. Knockout mice have decreased basal plasma glucose levels, and are resistant to insulin-stimulated glucose uptake in muscle and adipose tissue.
TBC1D4, Glucose, and Diabetes
As I mentioned, TBC1D4 has two different isoforms: a full-length protein, and a shorter isoform that’s missing exons 11 and 12. The p.R684X variant is in exon 11, so it likely only impacts the full-length isoform. Interestingly, that isoform is only expressed in skeletal muscle. Therefore it doesn’t affect TBC1D4 signaling in B-cells, the liver, or adipose tissue. In short, this variant’s disruption of the full-length protein causes higher plasma glucose levels, insulin resistance, and thereby confers risk of type II diabetes. It’s a sensible, straightforward story.
The work demonstrates that isolated founder populations are an important resource for mapping physiologically-relevant genetic variation associated with complex disease.
| Click to Tweet A new post on @MassGenomics on the value of isolated founder populations for genetic studies of complex disease: http://bit.ly/1lkJ3HH |

Thanks for pick up the story. I’ve read your blog for couple of months. As one of the co-author of the story, I also feel excited about the discovery. Just simple and beautiful.