This week marked an important milestone in our understanding of human genetic variation: the main publication of the 1,000 Genomes Project. The article in Nature describes the genomes from 1,092 individuals representing 14 populations across Europe, Africa, Asia, and the Americas. I think it’s important for anyone working in human genetics and genomics to read this paper, for a few reasons:
- It represents the most comprehensive characterization of rare variation, including SNPs, indels, and structural variants (SVs)
- The patterns of genetic variation reveal much about human population history and diversity
- The findings and methodology were produced by a collaboration included many (if not most) of the research leaders in sequencing, genomics, and human genetics.
The last reason may be one of the most significant achievements of this project, because it establishes (1) analysis methods and (2) a catalog of genetic variation that can leveraged by future studies. Indeed, participants in this project have driven forward advances in many areas of NGS analysis, including base quality recalibration, variant calling, and detection of structural variation.
Populations Sequenced
The 1,092 genomes sequenced comprise individuals from 14 populations, whose genomes were sequenced using a combination of exome and low-coverage whole-genome strategies. The populations are almost always referred to by their abbreviations, which are as follows:
- ASW, people with African ancestry in Southwest United States
- CEU, Utah residents with ancestry from Northern and Western Europe
- CHB, Han Chinese in Beijing, China
- CHS, Han Chinese South, China
- CLM, Colombians in Medellin, Colombia
- FIN, Finnish in Finland
- GBR, British from England and Scotland, UK
- IBS, Iberian populations in Spain
- LWK, Luhya in Webuye, Kenya
- JPT, Japanese in Tokyo, Japan
- MXL, people with Mexican ancestry in Los Angeles, California
- PUR, Puerto Ricans in Puerto Rico
- TSI, Toscani in Italia
- YRI, Yoruba in Ibadan, Nigeria
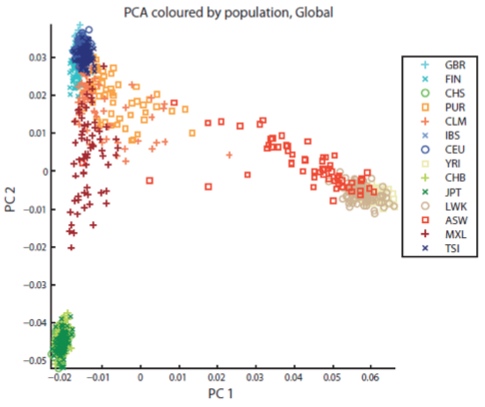
Though some are what we refer to as “admixed” populations, they all generally belong to one of four major groups of continental ancestry, and the members of each group tend to be related (as shown in the PCA plot above).
| Ancestry-based groups | ||
| AFR | African | YRI, LWK and ASW |
| AMR | Americas | MXL, CLM and PUR |
| EAS | East Asian | CHB, JPT and CHS |
| EUR | European | CEU, TSI, GBR, FIN and IBS |
Detection and Integration of SNPs, Indels, and SVs
A significant portion of the workload for this project was identifying an optimal set of variant calls for the sequencing data; I don’t envy the working groups who had to accommodate two, three, or even four methodologies for variant calling. Ultimately, however, the authors reached a consensus set of calls that establishes the new standard for variant detection in human genomes. In each individual, on average, they identified:
- 3.60 million single nucleotide polymorphisms (SNPs), of which 24,000 were in GENCODE (coding) regions
- 350,000 small indels (440 coding), confirming the expectation that these exist in a 1:10 ratio with SNPs in human genomes, and demonstrating the strong selection against indels in coding regions.
- 717 large deletions (the most confident category of SVs that we currently can detect), of which 39 overlapped GENCODE regions.
In the pilot phase of the project, the authors described the portion of the genome for which next-gen sequencing could provide informative variant detection as the “accessible” genome, which comprised 85% of its bases. Now, thanks to increases in read lengths and algorithmic improvements, that accessible portion has grown to include 95% of the genome. The remaining 5% is mostly low-complexity regions where accurate characterization of variants remains challenging.
Population Genetic Variation
The pilot phase of the 1,000 Genomes project and its predecessor the International HapMap Project had already identified and characterized common (MAF>5%) and less-common (MAF 1-5%) in the genome. The goal of the current study, in contrast, was to map rare variation present in less than 1% of human chromosomes. Such variation has been systematically under-represented in current studies of genetic variation, despite the fact that rare variants are likely to be enriched for functional changes. A comprehensive catalog of both rare and common variation, therefore, will provide a powerful resource for genome-wide association, Mendelian disease, and other human genetics studies.
In their 1,092 genomes representing 14 world populations, the authors found that:
- Most common variants (94%) with MAF>5% were known before the current phase of the project
- Variants present at MAF>10% overall were almost always present in all 14 populations
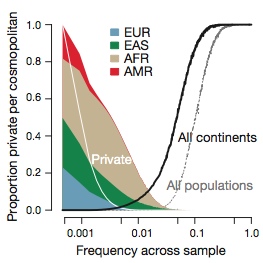
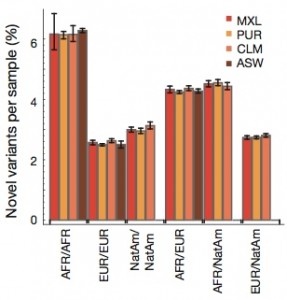
- The degree of rare-variant differentiation differed between populations. For example, FIN and IBS populations carry excesses of rare variants.
- Populations of African origin carry up to 3x as many rare variants as European or East Asian populations.
Functional Variants
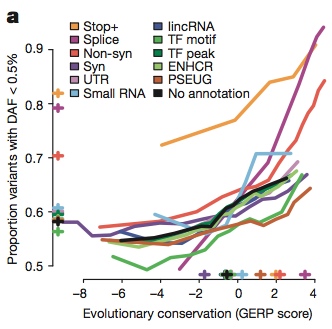
This study also represents the most comprehensive analysis of putative functional variation in healthy individuals. In essence, it’s a picture of the functional variants that most of us (without genetic diseases) are likely to harbor in our own genomes. The authors identified candidate functional variants using a few complementary strategies: gene annotation, experimentally-identified elements, and evolutionary conservation (GERP scores). For most types of variation, the observed level of purifying selection (a proxy for the functional importance of a variant) was correlated with conservation score:
Here, on the y-axis ou’re looking at the proportion of variants with derived allele frequency (DAF) of less than 0.5%… in other words, the fraction of variants in each class that showed very low variation since humans diverged from other primates. Higher on the y-axis suggests strong purifying selection against variants in a given category. And as we all know, purifying selection implies function.
You can note a couple of trends in the plot above. First, the strength of purifying selection trends nicely with evolutionary conservation, which is expected but also reassuring. Second, at least two categories (stop-gain, also called “nonsense” variants, and splice-site variants) exhibit dramatically higher levels of purifying selection than other classes, and with general disregard to conservation levels.
Imputation and GWAS
One immediate and powerful benefit to this dataset that we already saw with the HapMap and 1000 Genomes Pilot projects is a resource to aid imputation of missing genotypes in genetic association studies. Essentially, this boils down to linkage disequilibrium — the tendency of certain variants to be inherited together — and our ability to use that information to infer what a genotype is likely to be based on the genotypes that we do have. There are essentially two reasons you’d want to do this:
- To search for new signals of genetic association with a given phenotype
- To fine-map known associations, ideally to a single causative variant
Despite the different expected accuracies in calling intergenic SNPs versus exonic SNPs, small indels versus large deletions, the authors found that imputation accuracy was similar for these different types of variants. For low-frequency variants (MAF 1-5%), accuracy was 60-90% in all populations. That’s not bad, considering that imputation basically lets you get additional genotypes “for free”.
Fascinatingly, when the authors evaluated previous GWAS hits in Europeans, they found that each signal is, on average, in LD with 56 variants (51.5 SNPs and 4.5 indels). In 65% of such cases, there was at least one variant in LD with a high GERP score (>2) and 19% of the time, there was a coding variant in LD. This highlights two important facts about GWAS hits: they’re unlikely to be the causal variant themselves, but we can use resources like the 1,000 Genomes map to identify and follow-up on variants that could be functional.
Implications for Personal and Medical Genomics
The 1,000 Genomes project has provided a sort of “null expectation” for the number of rare, low-frequency, and common variants of different functional consequences found in randomly-chosen [healthy] individuals from various populations. It serves therefore as a kind of reference panel and benchmark for when we attempt to study individuals with some kind of phenotype — Mendelian disorders, cancer, disease susceptibility — to help pinpoint the differences. It also tells you that if you sequence an individual’s whole genomes and don’t find about 3 million SNPs, something is probably wrong.
So how many potentially deleterious variants do we expect to find in a given individual? The authors provide some rough estimates.
- 2500 nonsynonymous variants at conserved positions, of which 20-40 are likely to be damaging (2-5 of which are rare)
- 150 loss-of-function variants (splice site variants, stop gains, frameshift indels) of which 10-20 are rare
- 1-2 variants previously identified from cancer sequencing, which suggests either real somatic/acquired mutation, or (more likely), a small fraction of rare germline variants being submitted to the COSMIC database.
So it would seem that healthy individuals, at least, are on a somewhat level playing field: we all have some level of potentially deleterious variation in our genomes. Genes, environment, lifestyle, and numerous other factors (including luck) probably have an equal role in determining the health and well-being of any given person. That’s not surprising, if you ask me, and in fact, that’s kind of how you’d want it to be.
References
The 1000 Genomes Project Consortium (2012). An integrated map of genetic variation from 1,092 human genomes Nature DOI: 10.1038/nature11632

Neither Australia, south east Asia, Polynesian islands, the Indian sub-continent nor the middle east nor central Asia nor Russia nor eastern Europe are represented. While this is a great study, its not comprehensive and has great sampling bias. Can hardly be called substantial.
Thanks for the clear and very useful summary. One of the things I’ve been finding it hard to parse in the 1000 Genomes project is actually how many *complete* genomes have been sequenced to high coverage (say >30x) in humans. Is high coverage data from many individuals really only available for the exome (e.g. <5% of the genome)? How do you read the tea leaves on this?
Dan – great post – I’m sure a lot of people are grateful for the effort you put into these, many thanks.
Casey – the experimental design of the main 1000 Genomes project does not involve any high coverage whole genome samples. Two trios were sequenced to high coverage in the pilot, and there has been talk about doing more, but as far as this publication is concerned, there aren’t any (except I guess by chance when some samples have “accidentally” been over sequenced).
Feel free to push further questions directly to me if you want
regards
Zam