Cancer immunoediting is a process by which the immune system controls the growth of nascent tumors and shapes their antigenic properties. It’s a sort of catch-22 of cancer biology; by protecting the host from development of cancer, the immune system ultimately selects for tumor cells that are resistant to its attack. Central to the concept of immunoediting is the idea that T-cell recognition of tumor-specific antigens drives the destruction of early tumors, and later the antigenic “sculpting” of persistent tumor cells. And yet our understanding of the process of cancer immunoediting is incomplete, largely because most tumors that come under study are already highly edited, having arisen in hosts with competent immune systems. Two studies published online at Nature have shed light on the underpinnings of cancer immunoediting using carcinogen-induced or oncogene-driven models of cancer in genetically engineered, immune-deficient mice.
Cancer Exome Sequencing Reveals T-cell-dependent Immunoediting
Matsushita et al employed exome sequencing to carcinogen-induced sarcomas generated in Rag2-knockout mice, which lack both T-cells and B-cells. The parental tumors, called d42m1 and H31m1, are heterogeneous sarcomas likely comprised of several tumor sub-populations. Because they arise in immunodeficient hosts, they’re also highly immunogenic. When transplanted into wild-type mice with intact immune systems, these tumors are eliminated, presumably by T-cells. Using cDNA-exome capture and Illumina sequencing and the VarScan 2 algorithm, the authors performed systematic identification of somatic coding mutations in d42m1 and H31m1 tumors, finding:
- Largely distinct sets of non-synonymous mutations in each line (3,737 in d42m1; 2,677 in H31m1, and just 119 in common).
- Mutations in several cancer genes, including known lesions in Kras and Trp53 (mouse TP53) genes.
- A high proportion (46-47%) of C/A or G/T transversions, similar to the chemical-carcinogen signatures observed in human lung cancers.
- Mutation rates ten-fold higher than typical lung cancers in humans, though within three-fold of “hypermutator” tumors with mutations in DNA repair genes.
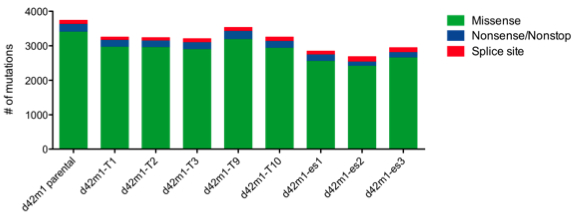
Interestingly, when parental d42m1 sarcoma cells were transplanted into wild-type mice, around 20% of recipients developed “escape” tumors which evaded immune destruction and progressed (escape clones). In contrast, the same cells transplanted to immunocompromised mice remained highly immunogenic; eight of ten are still eliminated when transplanted to wild-type mice (regressor clones). The remaining two, however, manage to grow in wild-type mice (progressor clones) with similar kinetics to the “escape” clones described above. Thus, the d42m1 sarcoma consists largely (but not entirely) of highly immunogenic clones and undergoes immunoediting in wild-type mice. Exome sequencing revealed that the parental lines (d42m1, H31m1), escape tumors (es1, es2, es3), eliminated clones (T1, T2, T9), and growth clones (T3, T10) all bore similar mutational loads:
Phylogenetic Analysis of Relatedness of Tumor Clones
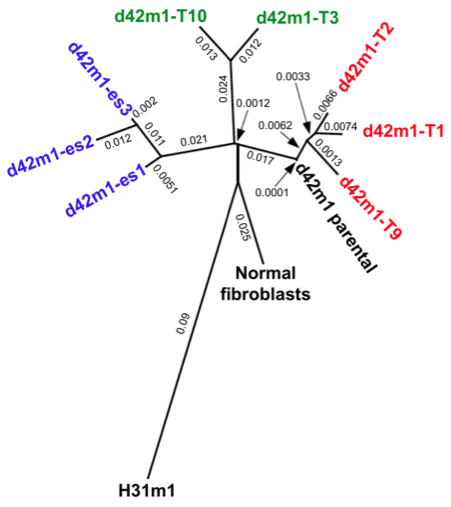
The systematic characterization of somatic mutations fed nicely into a phylogenetic analysis, which revealed that all d42m1 derived clones were related (genetically speaking) to one another, while quite distinct from the other sarcoma (H31m1) as well as normal fibroblast cells.
Notably, the regressor clones(T1, T2, T9) eliminated in wild-type mice were more closely related to the parental tumor, whereas the progressor clones (T3, T10) were more closely related to the escape tumors.
Searching for Tumor-specific Antigens
To investigate whether or not somatic coding mutations contributed to the different antigenicity observed between clones, the authors undertook an in silico analysis of predicted MHC class I binding affinity for each allele of missense mutations. Mutated proteins presented by MHC class I molecules are known to represent one class of tumor-specific antigens (TSAs) for CD8+ T-cells. In principle, somatic mutations affecting protein sequence might create epitopes that are presented on the surface of tumor cells by MHC class I, are recognized by T-cells, and cause an anti-tumor immune response.
The in silico analysis highlighted several candidate TSAs, but one was particularly striking: a missense mutation (p.R913L) in Spnb2, encoding the spectrin beta-2 protein. The mutant allele was expressed in d42m1 tumor cells and regressor clones, but absent from all escape tumors and regressor clones. To verify the importance of this mutation for d42m1 antigenicity, the authors performed conventional T-cell-based expression cloning. After three rounds of screening, they identified a single positive cDNA that was recognized by the C3 CTL clone, and its sequence was identical to the mutant Spnb2 protein.
T-cell-mediated Immunoediting Observed in vivo
It was clear that tumor cells lacking the mutant spectrin-B2 antigen represented only a minor subpopulation of the d42m1 tumor. This is apparent from the fact that most transplants of the tumor to wild-type recipients were eliminated. To determine if T-cell-mediated immunoselection was responsible for the outgrowth of the escape tumors, Matsushita et al performed a fascinating experiment using a mixture of cells from two d42m1 clones:
- d42m1-T2, a highly immunogenic clone eliminated by wild-type mice, labeled with red fluorescent protein
- d42m1-T3, a poorly immunogenic clone that progressed in wild-type mice, labeled with green fluorescent protein
The fluorescent labeling permitted the authors to monitor tumor outgrowth in vivo after mixtures were injected into either wild-type or Rag2-knockout (immune system deficient) mice. At a ratio of 95% T2 regressor cells and 5% T3 progressor cells, tumor outgrowth was observed in 5/5 (100%) Rag2-knockout recipients and 5/20 (25%) wild-type recipients. In essence, this re-capitulated the tumor outgrowth phenotype observed for the parental d42m1 tumor. If the authors depleted CD4+ or CD8+ T-cells in the recipients (using monoclonal antibodies) after challenge, 100% developed a tumor. Harvesting and assessing the outgrowth tumors in this experiment revealed that:
- Tumors from Rag2-knockout mice comprised 84% T2 cells and expressed mutant spectrin-B2, similar to the injected mixture.
- Tumors from wild-type mice comprised 98% T3 cells and did not expect mutant spectrin-B2.
Thus, in a single experiment, the authors demonstrated that (1) CD4+ and CD8+ T-cells were the mediators of immune system rejection of transplanted tumor cells, and (2) d42m1 escape tumors develop from T-cell-dependent selection favoring cells without the spectrin-B2 antigen. In other words, T-cell destruction of cells bearing mutant spectrin-B2 “sculpted” the tumor cells into poorly immunogenic cells lacking the antigen, which progressed and grew out.
See this WashU news article for more reading, and a video with senior author Robert Schreiber.
Expression of Tumor-Specific Antigens Underlies Cancer Immunoediting
Another study in the same issue of Nature also investigated cancer immunoediting in immunodeficient mice, this time in an oncogene-driven model of sarcoma. Michel DuPage and colleagues induced sarcoma through deletion of p53 and expression of oncogenic Kras (G12D) in Rag2-heterozygous (KP) or Rag2-knockout (KPR) mice. Some fancy genetic engineering allowed them to also deliver SIY and SIN antigens and thereby obtain immunogenic tumors. In a series of experiments, the authors demonstrated that:
- Lymphocyte-mediated protection from sarcoma formation requires CD8+ T-cells that respond to non-self antigens expressed in tumors.
- T-cells drive the immunoediting of cancer cells, selecting for those that either do not express potent antigens or are unable to present antigens to reactive T lymphocytes.
- Without carcinogens, it’s possible to generate sarcomas that lack potent tumor-specific antigens. In other words, tumor immunogenicity is not a universal characteristic of cancer development.
This is an elegant study that I hope to digest further. Go now, and read these papers!
References
Matsushita, H., Vesely, M., Koboldt, D., Rickert, C., Uppaluri, R., Magrini, V., Arthur, C., White, J., Chen, Y., Shea, L., Hundal, J., Wendl, M., Demeter, R., Wylie, T., Allison, J., Smyth, M., Old, L., Mardis, E., & Schreiber, R. (2012). Cancer exome analysis reveals a T-cell-dependent mechanism of cancer immunoediting Nature DOI: 10.1038/nature10755
DuPage, M., Mazumdar, C., Schmidt, L., Cheung, A., & Jacks, T. (2012). Expression of tumour-specific antigens underlies cancer immunoediting Nature DOI: 10.1038/nature10803
Great article and thanks for this nicely consolidated blog post! Your blog is really a nice tool–wish more sceintists were comfortable with social media…