A study now online at Cell employs has revealed new insights about the normal processes of aging and mutation, and their role in development of acute myeloid leukemia (AML).
Most tumors harbor numerous somatic mutations, but only a fraction are believed to contribute to cancer development and growth. In fact, most malignancies are thought to arise after a single initiating event, which may lead to genomic instability and thereby cause additional mutations. Through a process called clonal evolution, tumor cells acquire mutations and undergo natural selection for growth advantages.
Thus, many tumors (especially solid tumors) are not a uniform mass of identical cells, but a heterogeneous mixture of different cell subpopulations. They share the initiating mutation thanks to their common ancester, the “founding clone.” But they might have developed hundreds or thousands of mutations on their own. That’s one of the challenges faced by those of us who sequence cancer genomes. Among dozens of somatic coding mutations, which ones are the initiators, and which are just along for the ride?
Why Sequence AML?
AML offers an opportunity to study these processes because genomic instability is rare, and it’s believed that there are only a few initiating mutations. In this study, our group performed whole-genome sequencing on 24 AML cases. Twelve of these were FAB type M3, where the initiating event, a PML-RARA fusion, is known. The other twelve were FAB type M1 with normal cytogenetics, where the initiating event is usually unclear.
Mutations in AML
The success of this study in characterizing mutations in AML comes from two important components:
- Systematic validation by custom capture and deep sequencing of somatic SNVs, indels, and structural variation. This not only confirmed which predicted events were valid somatic mutations (an important step because these are rare), but provided deep read counts by which we could compute accurate allele frequencies and examine tumor clonality.
- Extension of the analysis to additional AML cases (53 M1 and 31 M3), in we screened for additional mutations in the 384 mutated genes. This provided a better picture of the prevalence of recurrent mutations in AML.
Among the 108 genomes assessed, there were 23 genes harboring non-silent mutations in at least three independent tumors, suggesting an important role in tumor development or growth. We observed an average of 14.5 “tier 1” (coding) mutations per tumor in the 24 WGS cases; on average, ~3 of these affected recurrently mutated genes in M1 tumors, and ~2 affected recurrently mutated genes in M3 tumors. This fits nicely with the idea that M1 genomes harbor an initiating mutation, a “driver”, analogous to the known driver (PML-RARA) in M3 genomes.
Driver Mutations in M1 Leukemia
Because the driver in M3 genomes is known, any mutations we find in M3 genomes aren’t likely to be initiating events. FLT3 mutations, for example, often co-occurred with PML-RARA in M3 genomes and are already known to be a cooperating (but not initiating) alterations. Therefore, we looked for genes recurrently mutated only in M1 genomes, and there were three:
- NPM1, a classic mutation known to be pathogenic in AML
- DNMT3A, encoding a DNA methyltransferase and found in the last couple of years to be recurrently mutated across various liquid tumors and even solid tumors
- IDH1, encoding isocitrate dehydrogenase 1, a gene with a “mutational hotspot” at amino acid 132, first observed in glioblastoma, and since found to be recurrent in numerous tumors including AML.
Background Mutations in Hematopoietic Stem Cells
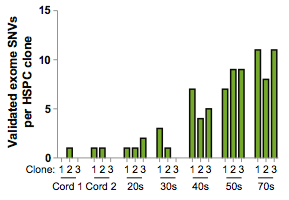
Now comes a second part of the study, something that sets it apart from other cancer sequencing efforts. We obtained human hematopoietic stem cells (the progenitors of blood cells) from 7 healthy volunteers of different ages (cord blood from newborns all the way to 70 year olds). For each sample, single HSCs were isolated and grown into colonies of homogenous cells to obtain enough DNA for exome sequencing. The exomes of each HSC colony were then compared to those of a matched blood normal, whose cells are produced by an estimated 1,000 operating HSCs. The idea is to determine, in healthy people, whether HSCs accumulate background mutations over time.
As it happens, yours truly had the task of analyzing HSC exomes and identifying somatic mutations. Here’s the thing about somatic mutation detection: the more mutations there are, the easier it gets. That’s why it’s possible to have a very high validation rate (>90%) for somatic mutations in solid tumors like lung cancer. There are just so many good, high-confidence mutations that the majority of them validate.
The HSC exomes, however, tended to have very few coding mutations (bad news for me, but good news for the healthy volunteers). Even with such tools as VarScan 2 and SomaticSniper, it was not an easy task. But we got it done, and the findings were striking: the number of coding mutations in HSC exomes directly correlated with the age of the volunteer. Cord blood cells from a newborn had virtually none, young people in that key disposable income marketing demographic had relatively few. Only over-the-hill volunteers in their 40’s and 50’s had appreciable numbers (5-9 mutations), and the 70-year-old had the most.
Mutations Correlate with Age in HSCs and AML
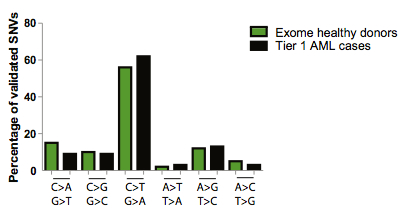
So mutations in the hematopoietic stem cells of healthy volunteers seemed to accumulate over time. Intriguingly, when AML patients were segregated by age, the correlation was also apparent. In other words, the number of mutations in both AML tumors and the stem cells from which they arise are correlated with age. Not only that, they’re similar in number and distribution across the mutation spectrum:
Background Mutation, Tumor Initiation, and Progression
This suggests that the majority of mutations in AML are random background mutations that occurred in HSCs. Then, an initiating mutation was acquired (e.g. NPM1 or PML-RARA). At that point, the background mutations in the cell were “captured” and carried forward as the progenitor gave rise to AML.
Cells from the founding clone sometimes acquired additional cooperating mutations (e.g. FLT3), yielding subclones that may have contributed to progression or relapse.
These findings reinforce many things that we already know: that mutations acquire gradually with age, that most of the mutations in AML (and likely other tumors) are random background events not contributing to tumorigenesis, and that subsequent mutation and evolution can give rise to subclones that ultimately determine cancer progression and response to therapy.
References
Welch JS, Ley TJ, Link DC, Miller CA, Larson DE, Koboldt DC, Wartman LD, Lamprecht TL, Liu F, Xia J, Kandoth C, Fulton RS, McLellan MD, Dooling DJ, Wallis JW, Chen K, Harris CC, Schmidt HK, Kalicki-Veizer JM, Lu C, Zhang Q, Lin L, O’Laughlin MD, McMichael JF, Delehaunty KD, Fulton LA, Magrini VJ, McGrath SD, Demeter RT, Vickery TL, Hundal J, Cook LL, Swift GW, Reed JP, Alldredge PA, Wylie TN, Walker JR, Watson MA, Heath SE, Shannon WD, Varghese N, Nagarajan R, Payton JE, Baty JD, Kulkarni S, Klco JM, Tomasson MH, Westervelt P, Walter MJ, Graubert TA, Dipersio JF, Ding L, Mardis ER, & Wilson RK (2012). The origin and evolution of mutations in acute myeloid leukemia. Cell, 150 (2), 264-78 PMID: 22817890