This month, Matthew J. Ellis and colleagues reported the whole-genome and/or exome sequencing of 77 estrogen-receptor-positive (ER+) breast cancer patients enrolled in a clinical trial of aromatase inhibitors. Their findings provide new insights into the genetic mechanisms of AI resistance, and may help pave the way to personalized cancer treatment in breast cancer.
Breast Cancer and Aromatase Inhibitors
Many breast cancers are “estrogen sensitive”, meaning that they need the hormone to grow. These tumors usually over-express the estrogen receptor (ER). ER-positive tumors comprise two “intrinsic subtypes”: Luminal A, which generally has a favorable prognosis, and Luminal B, which generally has a worse prognosis. Although ER-positive tumors represent the majority of breast cancer cases, they exhibit a wide range of prognoses, histological growth patterns, and treatment outcomes. Because of their dependence on estrogen, these tumors are often treated by estrogen deprivation therapy.
The well-known breast cancer drug Tamoxifen, for example, blocks a tumor’s ability to use estrogen. Aromatase inhibitors (AIs), in contrast, lower the amount of estrogen in the body by preventing non-ovary tissues from producing it. Some ER-positive tumors respond to AI therapy, and some do not. We don’t yet understand why this is. A clinical trial of AI therapy response paired with next-gen sequencing offers the opportunity to profile the genetic alterations of AI-sensitive and AI-resistant tumors.
Next-Gen Sequencing in a Clinical Trial
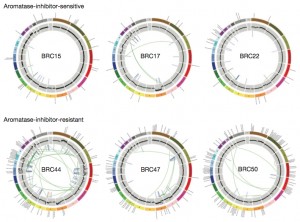
Seventy-seven cases from two neoadjuvant aromatase inhibitor clinical trials underwent next-gen sequencing (46 whole-genome, 31 exome). Of these, 29 were AI-resistant and 48 were AI-sensitive. The authors examined interactions between tumor proliferation levels (Ki67), histological categories, intrinsic subtype, and somatic alterations in these two categories of tumors.
As an author, I’m admittedly biased, but the sequencing and analysis in this study were state-of-the-art. Illumina paired-end whole-genome sequencing to 30x haploid coverage for tumor whole-genomes, and >80% coverage at 20x of CDS sequences for the exomes. Mutations were detected using SomaticSniper, VarScan, GATK, and Pindel. SVs were detected with BreakDancer and SquareDancer. High-level analyses were performed with our PathScan and MuSiC packages, as well as PathScan, GeneGo, MetaCore, and PARADIGM.
Candidate somatic mutations and SVs were experimentally validated by custom capture and deep resequencing. This not only enabled confirmation of the predicted events, but provided deep coverage to examine the clonal architecture of tumors. That’s critical when looking at therapy response. Further, the mutated genes were screened for recurrence in 233 additional breast cancer cases, for a total of 310 tumors surveyed.
Patterns of Somatic Alterations
This study also represents one of the largest surveys of breast cancer by next-gen sequencing, certainly by whole-genome sequencing. So what was the mutational landscape of luminal-type breast cancer?
- An overall mutation rate of 1.18 mutations per megabase (higher than AML, but lower than melanoma, liver, and lung cancers).
- The most frequently mutated gene was PIK3CA (41.3% of cases)
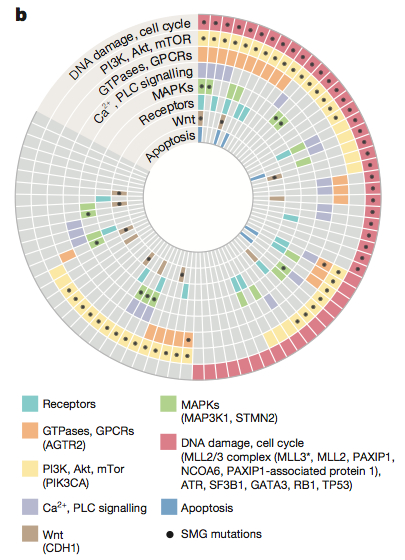
- 8 significantly mutated genes that are known breast cancer genes: PIK3CA, TP53, GATA3, CDH1, RB1, MLL3, MAP3K1, and CDKN1B
- 9 cancer genes not previously observed in clinical breast cancer samples: TBX3, RUNX1, MYH9, STMN2, SF3B1, CBFB, LDLRAP1, AGTR2, and STMN2
One of the relatively new findings is the recurrence of mutations in MAP3K1, a serine-threonine kinase that activates the ERK and JNK kinase pathways. Thirteen tumors had two non-silent MAP3K1 mutations (biallelic loss), and most of the mutations are highly deleterious (nonsense, frameshift, etc.) suggesting that this gene may act as a tumor suppressor. Across 310 cases, some 15.5% had mutations in MAP3K1 or MAP2K4.
Correlating Mutations with Clinical Data
A number of fancy clinical-correlation and pathway analyses revealed some interesting patterns of somatic mutations between AI-sensitive and AI-resistant tumors:
- TP53 mutations were significantly enriched in luminal B and high-grade tumors, and correlated with higher tumor proliferation (both at baseline and after AI therapy).
- MAP3K1 mutations, in contrast, were significantly enriched in luminal A tumors and correlated with lower proliferation at baseline.
- Several pathways were enriched in AI-resistant tumors, including TP53 signalling, DNA replication, and mismatch repair.
- ESR1 and FOXA1 were among activated hubs in the entire cohort, while MYC, FOXM1, and MYB were activated in AI-resistant tumors.
- GATA3 mutations were not associated with Ki67 levels, but did correlate with reduced Ki67 over therapy, suggesting it may be a positive predictive marker for response to aromatase inhibition.
In summary, this study sheds light on the somatic alteration landscape of ER-positive breast cancers and offers insight into some of the mechanisms of aromatase inhibitor resistance.
References
Ellis MJ, Ding L, Shen D, Luo J, Suman VJ, Wallis JW, Van Tine BA, Hoog J, Goiffon RJ, Goldstein TC, Ng S, Lin L, Crowder R, Snider J, Ballman K, Weber J, Chen K, Koboldt DC, Kandoth C, Schierding WS, McMichael JF, Miller CA, Lu C, Harris CC, McLellan MD, Wendl MC, DeSchryver K, Allred DC, Esserman L, Unzeitig G, Margenthaler J, Babiera GV, Marcom PK, Guenther JM, Leitch M, Hunt K, Olson J, Tao Y, Maher CA, Fulton LL, Fulton RS, Harrison M, Oberkfell B, Du F, Demeter R, Vickery TL, Elhammali A, Piwnica-Worms H, McDonald S, Watson M, Dooling DJ, Ota D, Chang LW, Bose R, Ley TJ, Piwnica-Worms D, Stuart JM, Wilson RK, & Mardis ER (2012). Whole-genome analysis informs breast cancer response to aromatase inhibition. Nature, 486 (7403), 353-60 PMID: 22722193