On the flight to Marco Island I took a stack of papers from Genome Research’s Cancer Issue. One of these is a study of epigenetics in breast cancer by Gary C. Hon and colleagues from the Ludwig Institute for Cancer Research. Hands down, this is one of the best papers I’ve read this year – interesting subject matter, elegant experiments, and beautiful presentation (in the form of several color figures). Using cutting-edge techniques enabled by next-generation sequencing, the authors generated complete methylome maps at single nucleotide resolution in a low-passage breast cancer cell line and normal breast tissue (primary human mammary epithelial cells).
A clear and growing body of evidence supports the notion that epigenetic alterations, such as DNA methylation, occur early in and contribute to tumor formation. In breast cancer, several known cancer genes, such as BRCA1 and CDH1 (E-cadherin) are commonly silenced by hypermethylation. Recent genome-wide assays for DNA methylation have reported that global hypomethylation is a hallmark of breast and colon cancers. Such epigenetic mechanisms have already been exploited for diagnosis and treatment; for example, DNA hypomethylation therapy with 5-azacytidine is approved for MDS and is in clinical trials for some other cancers.
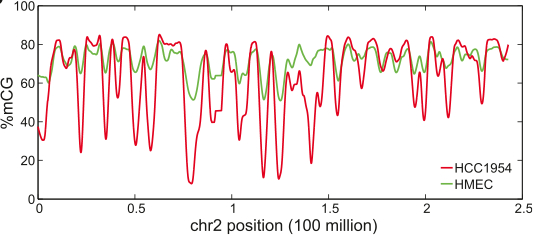
Hon et al created a low-passage cancer cell line using a Her2+ ER-/PR- tumor from an East Indian female. As a normal comparator, they obtained primary epithelial mammary cells. Both samples were assessed by whole-genome shotgun sequencing (~27x coverage) of bisulfite-treated DNA, or MethylC-seq, to assess DNA methylation:
This unbiased approach revealed global hypomethylation in the cancer cell line (HCC1954) relative to epithelial cells (HMEC), and some specific patterns:
- Methylation occurred almost exclusively (>99.8%) in the CG context
- Domains of hypomethylation spanned several known tumor suppressor genes such as DACH1.
- Loci with the most pronounced methylation changes in HCC1954 coincide with regions not fully methylated in epithelial cells.
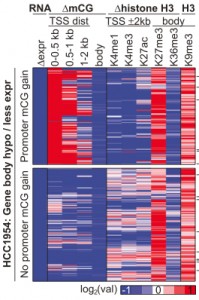
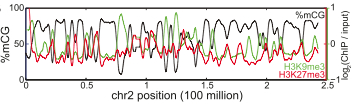
Repressive histones (red) increased in regions of hypomethylation. Hon et al, Genome Res (2012). Figure 2D.
This last observation suggests that certain partially-methylated regions of the genome consistently gain or lose methylation in cancer.
The dogma of DNA methylation is that it generally represses gene expression – in other words, methylated regions are transcriptionally silent, and unmethylated regions are often transcriptionally active. Here, the authors encountered a puzzling result. RNA-seq analysis of the tumor cell line revealed a number of silenced genes (presumably including many tumor suppressors). Integrating these with methylation data revealed that half fell into an expected pattern: hypermethylated promoters which repressed gene expression. The other 54%, however, were hypomethylated in the cancer cell line. The genes were silenced, but it wasn’t from methylation. The authors looked at gene expression in a panel of 50 ERBB2 (Her2) positive breast tumors and found that this pattern held up: DNA hypomethylation is associated with decreased gene expression in breast cancer.
Active and Repressive Chromatin by Histone CHiP-Seq
Repressive histone marks were another possibility. To evaluate histone modification (another epigenetic event), the authors employed chromatin immunoprecipitation with high-throughput sequencing (CHiP-Seq) to capture DNA sequences bound by H3 histones marking active (K4me1, K4me3, K27ac, K36me3) or repressed (K27me3, K9me3) chromatin. Plotted together with the DNA methylation from MethylC-seq, the integrated picture of epigenetic modifications was quite striking:
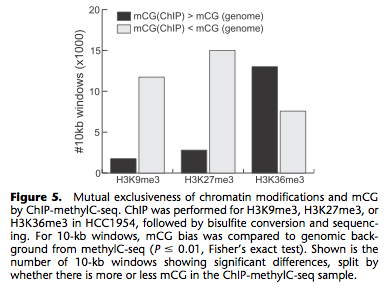
In regions of DNA hypomethylation, H3K27me3 and H3K9me3 form repressive chromatin domains. Thus, another form of epigenetics provided compensatory gene silencing. Further work revealed that DNA methylation and K27/K9 repression were mutually exclusive:
Haplotype Inference for Allele-Specific Epigenetics
Next, the authors wanted to investigate the allelic basis of DNA methylation and repressive chromatin. They used WGS data and the bam2mpg genotyping tool to call 1.2 million high-confidence SNPs genome-wide. These were error-corrected and utilized to infer haplotypes using HASH and HAPCUT, yielding 269,392 phased haplotype blocks. Integrated with the epigenetic data, these haplotypes revealed that:
- Almost half of partially-methylated (40-60%) blocks exhibited allele-specific methylation.
- 78% of H3K36me3 (activating) histone blocks were on the same allele as methylated (mCG) regions.
- In contrast, the majority of repressive histones H3K9me3 (75%) and H3K27me3 (79%) are on the opposite allele as mCG.
High-Level Organization of Repressive Histones
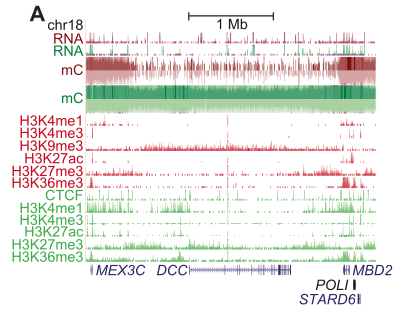
Finally, and this is very cool, the authors observed high-level organization of the regions repressed by H3K27me3 and H3K9me3, visualized here across the region of the DCC tumor suppressor gene on chromosome 18:
Large regions bound by H3K9me3 are flanked by smaller regions bound by H3K27me3, forming a histone “repressive domain” that silences gene expression. These two histones thus have a key role of epigenetic silencing of tumor suppressor genes in breast cancer (and likely other cancers), suggesting that they may represent new therapeutic targets. Further, the compensatory mechanisms of DNA methylation and histone repression suggest that targeting both alterations will probably be required to achieve successful therapy.
Go read this paper. It’s pretty.
References
Hon, G., Hawkins, R., Caballero, O., Lo, C., Lister, R., Pelizzola, M., Valsesia, A., Ye, Z., Kuan, S., Edsall, L., Camargo, A., Stevenson, B., Ecker, J., Bafna, V., Strausberg, R., Simpson, A., & Ren, B. (2011). Global DNA hypomethylation coupled to repressive chromatin domain formation and gene silencing in breast cancer Genome Research, 22 (2), 246-258 DOI: 10.1101/gr.125872.111