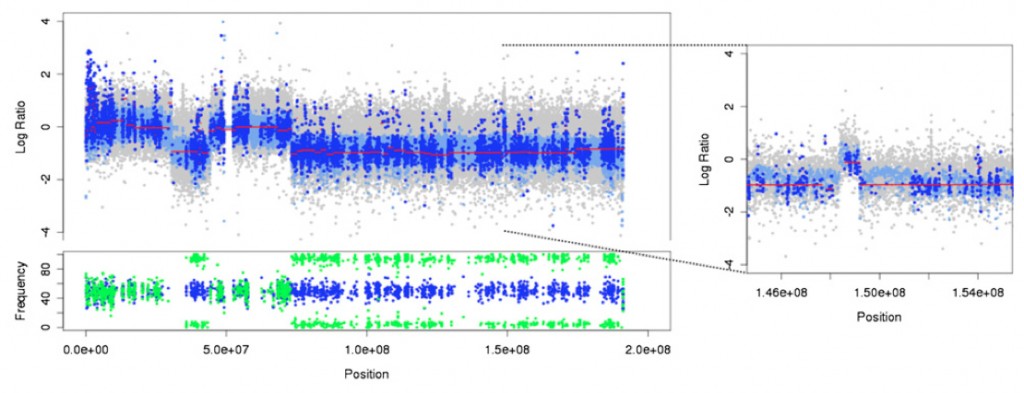
Now online at Genome Research is the publication of VarScan 2, our in-house algorithm for simultaneous detection of somatic mutations and copy number alterations using exome sequence data from matched tumor-normal pairs. There are a number of reasons why exome-based copy number alteration (CNA) detection should not work. The hybridization process introduces biases, both between samples and between targeted regions. And yet, by sequencing tumor samples and matched (closely related) normals under identical protocols, we demonstrated that read-depth-based somatic CNA detection can be performed with astonishing accuracy. Take this example:
Looking at a single chromosome (in the above case, chromosome 4 from an ovarian tumor), several patterns are immediately apparent. Obviously SNP array (grey) and whole-genome sequencing have denser coverage of the chromosome as a whole. The exome, however, appears fairly extensive. This is in part due to the nature of hybridization capture – there’s a shouldering effect in which sheared fragments in the genomic library that partially overlap target probes are captured, giving you coverage for 50 bp, 200 bp , or even 500 bp to either side of the exon. With paired-end reads and standard library size, every targeted exon generates coverage across at least 250 bp no matter its size. Another factor is off-target coverage, the regions that you didn’t enrich for but got anyway. While this non-coding sequence space is less valuable for finding protein-altering mutations, it’s still informative for copy number changes (amplifications and deletions), since these tend to be larger.
Thus, although current exome kits target 40-60 Mbp of the genome, they provide some coverage of nearly twice that.
Copy Number Alteration Detection Algorithm
The new algorithm in VarScan (“copynumber”) is very straightforward. Reading SAMtools “pileup” or “mpileup” input for normal and tumor samples simultaneously, it examines each position where both samples met the user-specified minimum coverage threshold, say 8x. These positions are grouped into contiguous regions of coverage, whose boundaries are defined by one of the following:
- A position that does not meet the minimum coverage requirement
- A non-covered position, observed as a position distance > 1 in the pileup.
- The start or end of the current chromosome, or
- A significant change in the regional log2 ratio of tumor-to-normal depth.
The output of the copynumber algorithm is thus a set of regions with chromosome, start, stop, normal depth, tumor depth, and a log2 ratio of tumor/normal representing the relative copy number change. In essence, this is identical to array-based copy number data, and therefore amenable to the same segmentation algorithms. We applied circular binary segmentation (CBS) using the “DNAcopy” R library of the Bioconductor package to segment the raw regions and identify significant change-points. The output of CBS is what’s shown above.
Comparison to SNP Array and Whole Genome Sequencing Data
In the paper, we compared exome-based copy number results to SNP array and whole-genome sequencing copy number results (the latter segmented into 10kbp bins and called by cnvHMM) for five ovarian cancer cases. The amount of overlap was surprising, particularly for large-scale events encompassing >25% of a chromosome arm:
Our exome-based approach detected 88% of large-scale events identified by array or WGS, and 96% of events detected by both array and WGS. Large-scale events not called were visible in the exome results, but generally were over-segmented into smaller events on chromosome arms with sparse exome coverage. Thus, exome-based copy number analysis with VarScan 2 detects the vast majority of large-scale copy number alterations despite the targeted nature of the data.
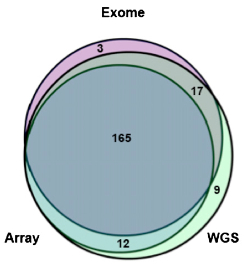
Next, we looked at focal events detected by the three platforms. This is a harder comparison to make, since SNP array and exome both target specific regions of the genome, while only WGS provides an unbiased survey. So, we focused on somatic CNAs affecting genes, which should theoretically be detectable by all three platforms:
Here, the results may be even more exciting. A substantial fraction of events were detected by all three platforms, but each also contributed a set of unique focal CNAs. More than 80% of the exome-based CNAs were supported by at least one other platform, suggesting that they’re likely to be real events. Yet the exome results contained a high fraction of platform-specific calls, possibly focal genic CNAs that were missed by the other two platforms. In other words, exome-based copy number analysis may detect small CNAs of coding regions missed by conventional platforms.
Applications of Exome-based Copy Number Analysis
The ability to detect somatic CNAs with high precision using exome data has enormous potential because there are so many exome datasets out there amenable to it. Literally thousands of tumors have undergone exome sequencing in the past few years; many of them were not evaluated for somatic copy number changes or were assessed by array-based methods that obviously miss things. Detecting large-scale and focal CNAs in addition to somatic mutations and germline variants in a single package (VarScan 2) may therefore offer a fairly comprehensive view of genetic variation in the coding regions of tumor genomes.
References
Koboldt DC, Zhang Q, Larson DE, Shen D, McLellan MD, Lin L, Miller CA, Mardis ER, Ding L, & Wilson RK (2012). VarScan 2: Somatic mutation and copy number alteration discovery in cancer by exome sequencing. Genome research PMID: 22300766