Contents: AML Sequencing • Capture Validation • Somatic Alterations • Mutation Profile • Evolution and Clonality • Convergent on IDH2 • Carl Zimmer Article • References
Acute myeloid leukemia (AML) is a cancer of myeloid blood cells, in which abnormal white blood cells accumulate in the bone marrow and interfere with normal blood cell production. This is a highly malignant tumor affecting 13,000 adults in the United States each year; if left untreated, it progresses rapidly and leads to death within weeks or months. The standard treatment is chemotherapy: induction therapy to achieve remission, followed by consolidation therapy to eliminate any residual disease. Most of the 8,800 annual deaths in the United States are of patients who relapse with a tumor that has undergone clonal evolution at the cytogenetic level. These relapsed tumors, unlike primary AML, are resistant to chemotherapy and progress rapidly.
Whole-genome sequencing of Relapsed AML
By sequencing the complete genomes of primary tumor, relapsed tumor, and matched normal (skin) samples from 8 AML patients, our group was able to study clonal evolution of AML tumors at the genetic level. The patients comprised 5 different French-American-British hematological subtypes; the time to relapse after initial diagnosis ranged from 235 to 961 days. One of these 8 cases was from patient UPN 933124, which we informally call “AML1” and whose primary tumor was the first cancer genome to be published, also by our group, back in 2008. Whole-genome sequencing of the 8 cases (24 samples) achieved >25x haploid coverage with >97% of diploid alleles represented.
Custom capture and deep sequencing validation
One technical achievement of this project was the high-throughput, deep sequencing validation strategy. For each patient, we designed a custom hybridization capture array (Nimblegen) targeting all candidate somatic events from the primary tumor and relapse sample (median: 539 per case). This enabled genome-wide validation of all variants in uniquely mapped regions, providing a sufficiently large set of data points for clonality analyses. Such a strategy is especially critical in AML, as tumors harbor relatively few somatic coding mutations (~21 per tumor in this study; the range is typically 5 to 50). Fragmented DNA from the primary tumor, relapse sample, and skin (normal) sample was individually hybridized with the validation probeset. Captured libraries then underwent deep sequencing on the Illumina platform, achieving a median of 590-fold coverage for each site. Such high redundancy of coverage enabled us to accurately compute allele frequency, and thereby, the fraction of tumor cells harboring each mutation.
Patterns of somatic alteration
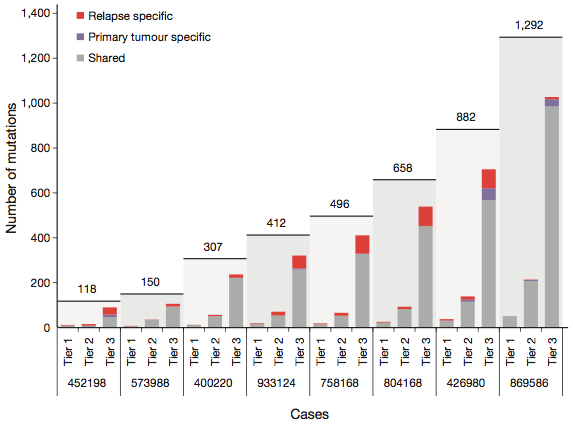
We validated a total of 4,315 somatic events genome-wide. These included tier 1 (coding), tier 2 (conserved/regulatory), and tier 3 (unique noncoding) variants.
As expected, tier 1 mutations comprised the smallest category and tier 3 the largest in all tumors. This generally reflects the proportion of the genome in each tier (tier 1 is just 5%), though we and other groups have observed that the mutation rates of many tumors are lower in coding regions, likely due in part to transcription-coupled DNA repair. We utilized exome data for 200 AML cases sequenced by the Cancer Genome Atlas research network to identify recurrently mutated genes. These included:
- Known AML genes, including DNMT3A, FLT3, NPM1, IDH1, IDH2, WT1, RUNX1, PTPRT, PHF6, and ETV6
- Novel recurrently-mutated genes, including WAC, SMC3, DIS3, DDX41, and DAXX.
Details on the recurrently mutated genes, and structural and functional analysis of somatic rearrangements, are provided as supplementary materials.
Mutational Profile of Relapsed AML
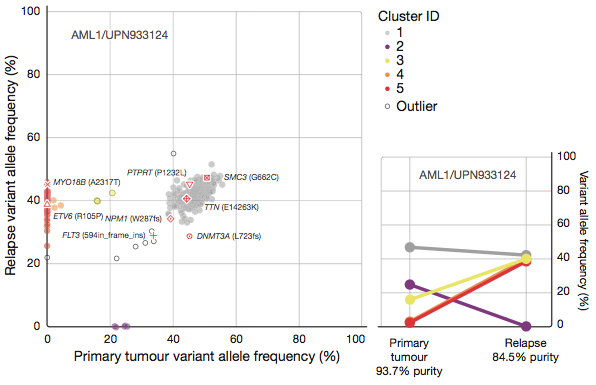
AML1 (UPN 933124) exemplifies the analysis approach we applied to relapsed AML. There were 413 validated somatic events in AML1, of which 78 were relapse-specific, 5 were primary-tumor-specific, and 330 were shared between tumors. Deep sequencing validation revealed some interesting allele frequency patterns for validated mutations:
You will note that most of the somatic events found in the primary tumor were also present in the relapse and vice-versa. Assuming that all mutations are heterozygous (which is likely), the observed allele frequencies suggest that shared mutations are present in virtually all tumor cells. The allele frequencies are higher in the primary tumor because its tumor cellularity, or purity (93.7%) was higher than that of the relapse (84.5%). Notably, we estimated that the tumor content of the “normal” skin sample was 29% due to infiltrating leukemic cells. This illustrates a key challenge in studying leukemia: most of the somatic mutations are observed at moderate allele frequency in the matched normal. A simplistic approach to somatic mutation detection, in which one simply subtracts all variants called in the normal from those called in the tumor, is poorly suited here. VarScan 2 and SomaticSniper are two mutation-detection algorithms developed by our group capable of addressing this problem.
Tumor Clonality and Evolution
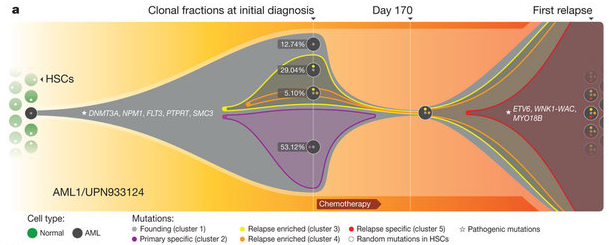
A clustering analysis of mutant allele frequencies suggested that there were four clones (tumor subpopulations) in the primary tumor defined by distinct sets of mutations.
- Clone 1 (46.86% of primary tumor) was the founding clone.
- Clone 2 (24.89% of primary tumor) was derived from clone 1.
- Clone 3 (16.00% of primary tumor) was derived from clone 1.
- Clone 4 (2.39% of primary tumor) likely arose from clone 3.
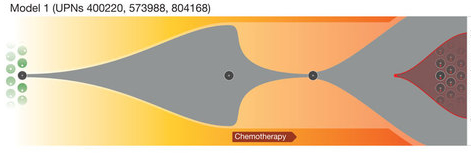
The evidence suggests that a relatively minor subpopulation of tumor cells (clone 4) survived chemotherapy and arose to become the dominant clone at relapse. In the process, it gained additional mutations, possibly via the DNA damage induced by chemotherapy. Four other AML cases in this study were consistent with this model of tumor evolution. We also observed another model, in which the dominant clone in the primary tumor gained mutations found only at relapse:
Three of the cases in this study were consistent with this model of tumor evolution.
Convergent Evolution of IDH2 Mutations
Here’s an interesting side-story to this study that wasn’t really discussed in the paper. Two cases harbored mutations in isocitrate dehydrogenase 2 (IDH2), a gene known to be recurrently mutated in AML, glioblastoma, and other tumors. In patient AML28 (UPN 573988), we detected a C to A substitution at chr15:88432938 (build36) causing an amino acid change (R140L) in IDH2. This mutation was also present in the relapse sample at a moderate frequency. However, we also detected a G to A mutation at chr15:88432938 (one base downstream) affecting the same arginine residue but changing it to tryptophan (R140W) instead of leucine. And these two mutations were mutually exclusive. Thus, two subclones of the tumor in AML35 both acquired activating mutations of IDH2 at the same residue but by different mutations.
Zimmer Article on Discover Blogs
Carl Zimmer has a nice article on this study on his blog at Go now, and read this paper.
References
Ding, L., Ley, T., Larson, D., Miller, C., Koboldt, D., Welch, J., Ritchey, J., Young, M., Lamprecht, T., McLellan, M., McMichael, J., Wallis, J., Lu, C., Shen, D., Harris, C., Dooling, D., Fulton, R., Fulton, L., Chen, K., Schmidt, H., Kalicki-Veizer, J., Magrini, V., Cook, L., McGrath, S., Vickery, T., Wendl, M., Heath, S., Watson, M., Link, D., Tomasson, M., Shannon, W., Payton, J., Kulkarni, S., Westervelt, P., Walter, M., Graubert, T., Mardis, E., Wilson, R., & DiPersio, J. (2012). Clonal evolution in relapsed acute myeloid leukaemia revealed by whole-genome sequencing Nature DOI: 10.1038/nature10738